Locus Rank
20Gene
Id
256Gene Name
NXPH4Duplicated
FalseMaternal
FalseGene Not Within Locus (Nearby
FalseGene Description
Mouse Phenotype
decreased circulating triglyceride level/increased urine nitrite levelAdditional Image
Allen Brain
Allen Regions
Zfin In Situ
Zfin Brain Areas
Published Zebrafish Pehnotype
Not In Allen Brain
FalseZFIN Link
http://zfin.org/ZDB-GENE-141017-1Allen Link
http://mouse.brain-map.org/gene/show/68246More Additional Images
Papers
Locus Rank
Locus Rank
20Genes in Region
LRP1 MYO1A NAB2 NDUFA4L2 NXPH4 R3HDM2 SHMT2 STAC3 STAT6 TAC3 TMEM194A
Associated Snps
rs12826178 (21)
rs324017 (103)
GWAS Region (hg19)
chr12:57428314-57682971Genes Skipping
MYO1A - no allen brain, myosin; NDUFA4L2 - no allen brain, nadh dehydrogenase; NXPH4 - high in hindbrain of allen and jax knockout doesn't sound promising . . .; STAC3 - muscle (there is one 2015 paper hinting at calcium channels in brain, but zfin shows muscle expression only); TAC3 - is neuro, but a lot is known and seems like it's not this gene (has to do with puberty though . . ); TMEM194A - nothing is known, but not in allen brainRicopili Plot Surrounding Area
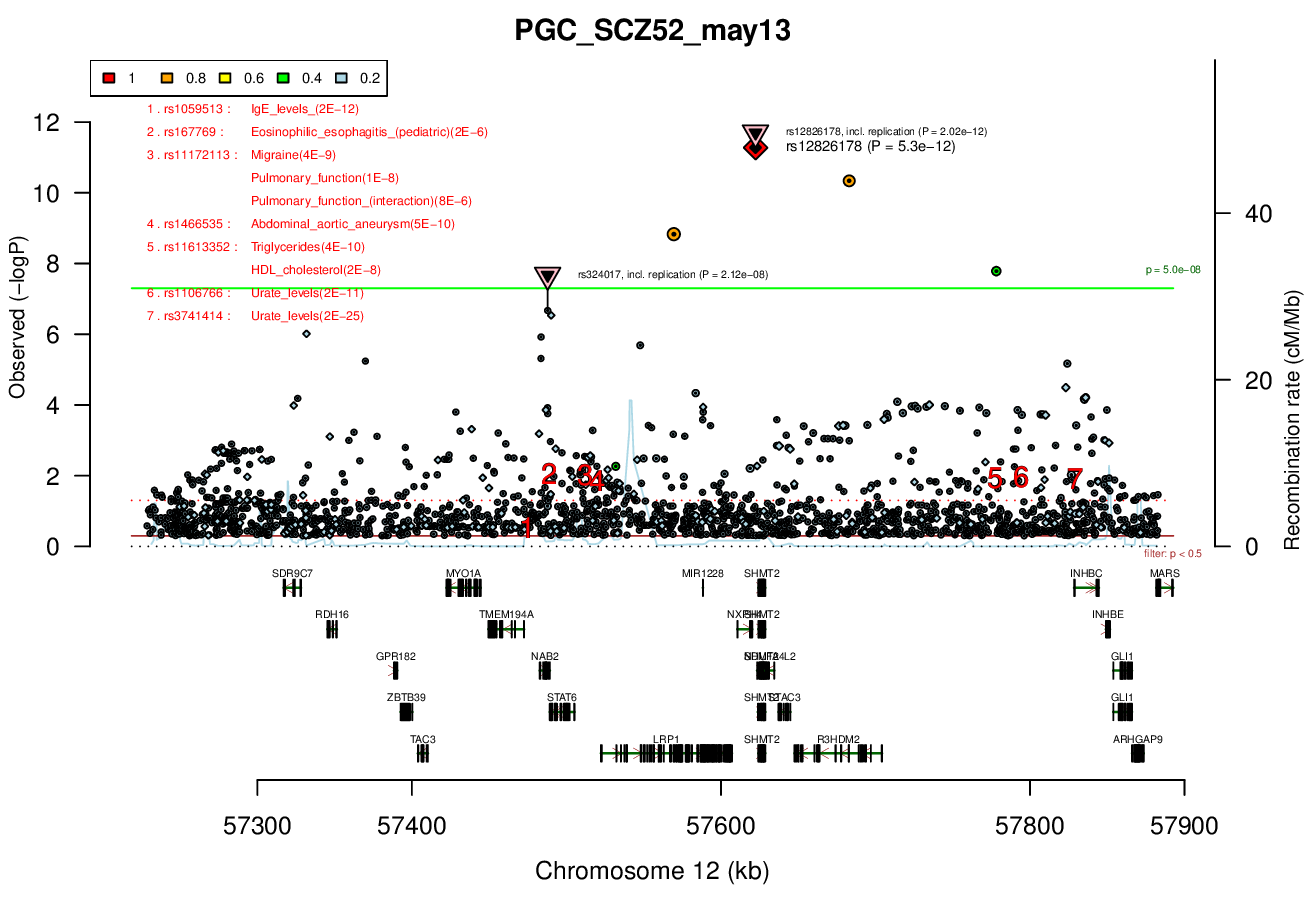

Distance On Each Side Of Locus In Above Plot
0.2 MBRicopili Plot
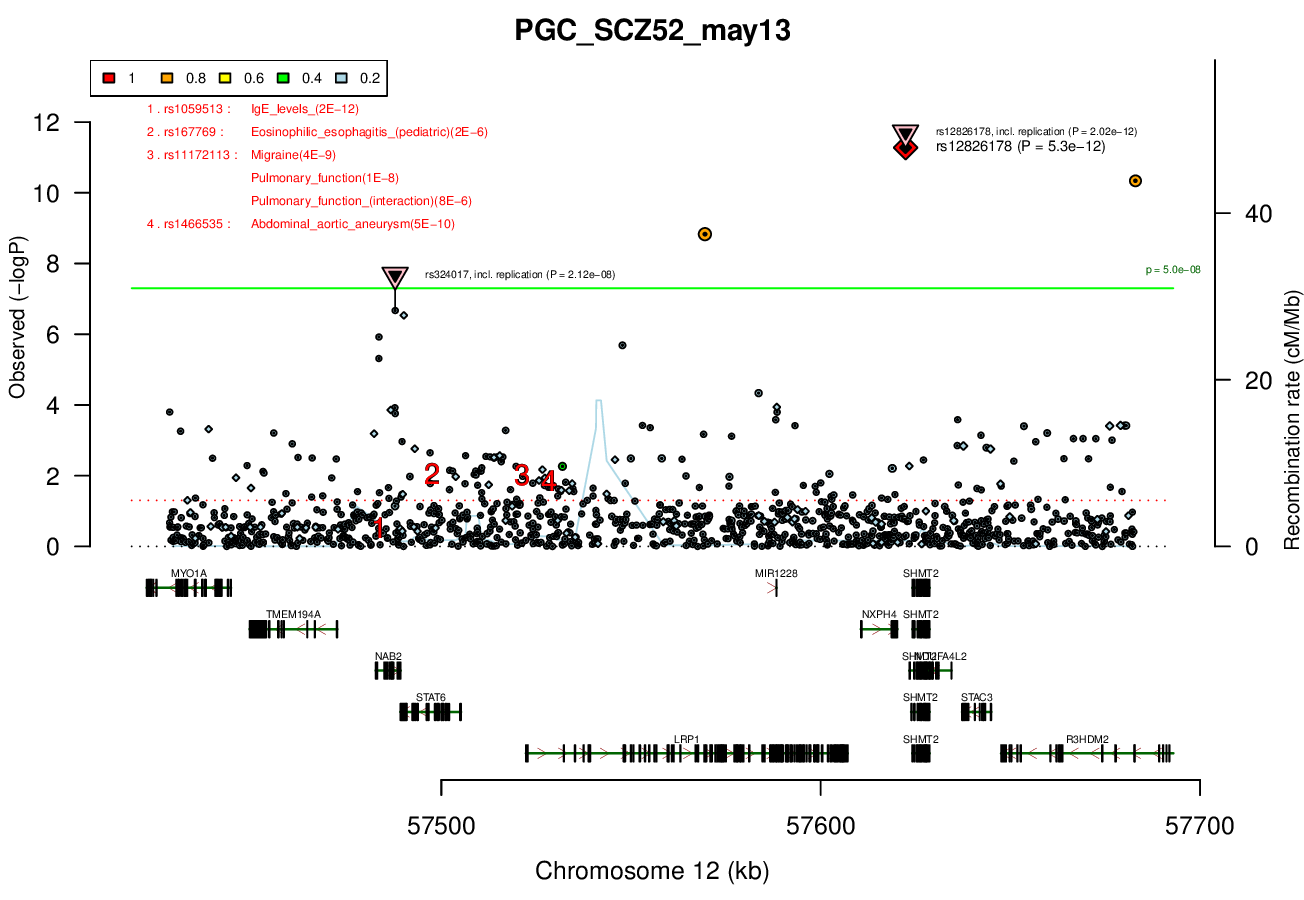
Gwas And Surrounding Region Pic
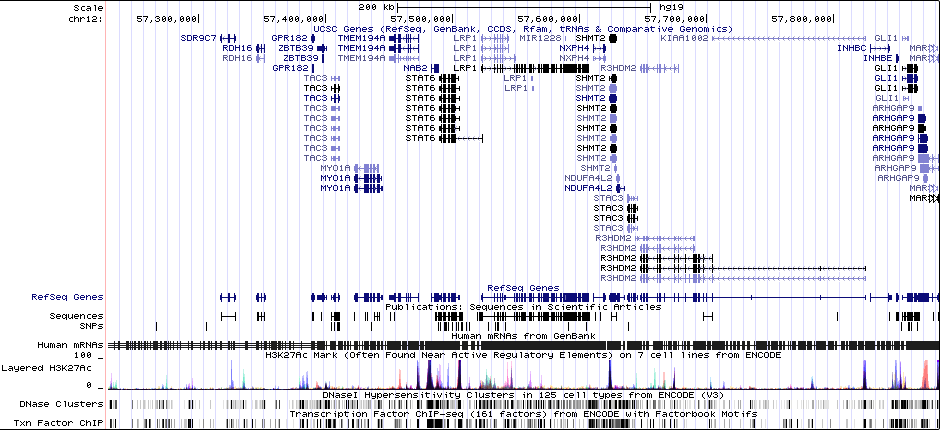
Gwas Region Pic
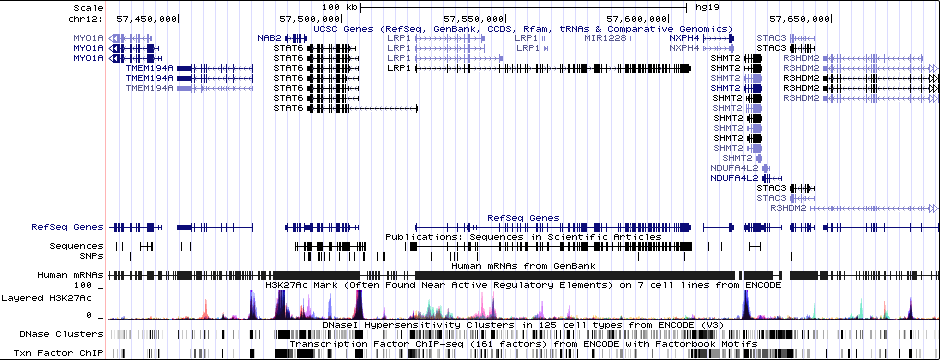
Sequence 1
Ensembl Gene Name
NXPH4Harvard Allele
a309Mutation Area - WT DNA Sequence
ggtCTCTAGATTATGTGGATCCGGGATCAACAGGTAGCGTTTTCAAGAGCGTCCCGTACGGCGGACTGAAGCCCCCCACACACACCTCCCGAATATTCCCCCCTGTCGACCACAAGCCCAAACCACCACCACCCGGCTTCAGCTTCCACAACCCGTACGAGTGGGCACGCAACCAGACGCACCTGCCGGACCAGAGTGGCTACCGGCCCAAACGCAAGCCCACCCTCAAGACCTCCATGAAGACCAAGAAGATCTTCGGCTGGGGCGACTTCTACMutation Area - Mutant DNA Sequence
GGTCTCTAGATTATGTGGATCCGGGATCAACAGGTAGCGTTTTCAAGAGCGTCCCGTACGGCGGACTGAAGCCCCCCACTTCAGCTTCCACAACCCGTACGAGTGGGCACGCAACCAGACGCACCTGCCGGACCAGAGTGGCTACCGGCCCAAACGCAAGCCCACCCTCAAGACCTCCATGAAGACCAAGAAGATCTTCGGCTGGGGCGACTTCTACGuide RNA Target Sites
TTCAAGAGCGTCCCGTACGGCGG
ATATTCGGGAGGTGTGTGTGGGG
TTGTGGAAGCTGAAGCCGGGTGG
TGGTCTTCATGGAGGTCTTGAGG
Genotyping Primers
f, ggtCTCTAGATTATGTGGATCCGG
r, GTAGAAGTCGCCCCAGCCG
Alignment of WT and Mutant Sequences w/ gRNA Targets (cyan) and Genotyping Primers (pink) Shown on WT Sequence
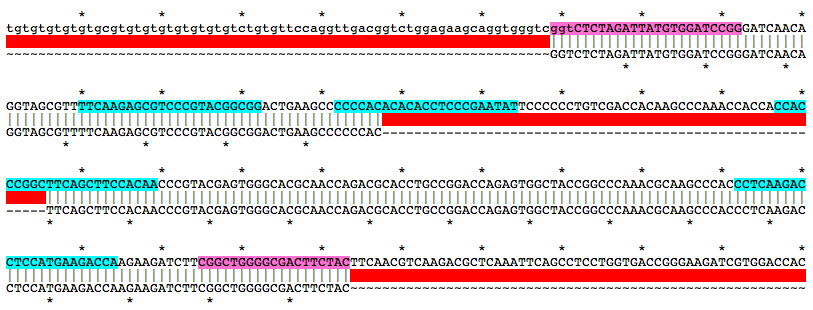
Allele
58bpDWT Genotyping Size
275WT Protein Sequence
MRLLCWSFLLLIQWILRKVDGLEKQVGRSLDYVDPGSTGSVFKSVPYGGLKPPTHTSRIF
PPVDHKPKPPPPGFSFHNPYEWARNQTHLPDQSGYRPKRKPTLKTSMKTKKIFGWGDFYF
NVKTLKFSLLVTGKIVDHINGTFTVYFRHNSSSLGNVSVSIVPPTKIVEFEIQNPLHPEI
QFHPPQQSTIDPKDIKTFNCRVEYEKTDRSKKPKPCLYDPSQTCFTENTQSHSSWLCAKP
FKVICIFISFFSIDYKLVQKVCPDYNFQSEHPYFG-
Mutant Protein Sequence
MRLLCWSFLLLIQWILRKVDGLEKQVGRSLDYVDPGSTGSVFKSVPYGGLKPPTSASTT
RTSGHATRRTCRTRVATGPNASPPSRPP-
Mutant Genotyping Size
217Behavior Data
Behavior Data Summary
NoneBehavior Data Description 1
Dark flash block 1 start Merged section Pvalue = not significant 10 het vs 10 hom ribgraph_mean_ribbon_fullboutdata_dpix_a0darkflash103.pngBehavior Data Graph 1
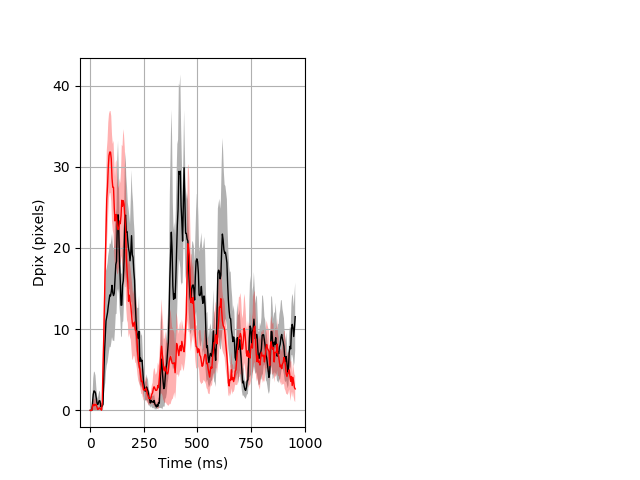
Behavior Data Description 2
Day all prepulse tap Merged section Pvalue = 0.0337 10 het vs 10 hom ribgraph_mean_ribbon_fullboutdata_dpix_dayprepulseinhibition100d.pngBehavior Data Graph 2
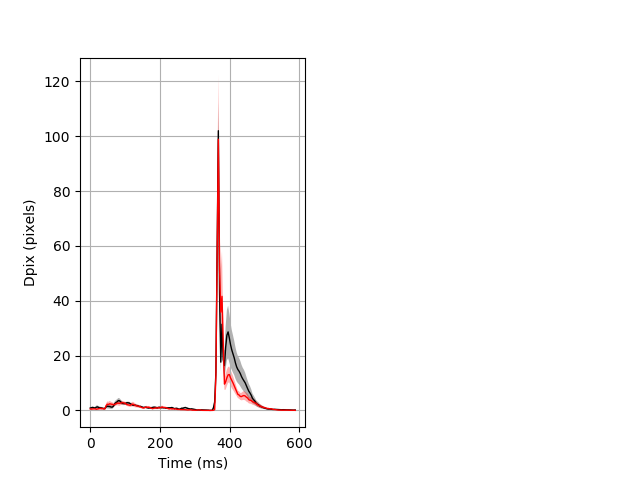
Behavior Data Description 3
Features of movement entire protocol Graph Pvalue = not significant Merged section Pvalue = not significant 10 het vs 10 hom ribgraph_mean_ribbonbout_aveboutdisp_10min_combo.pngBehavior Data Graph 3
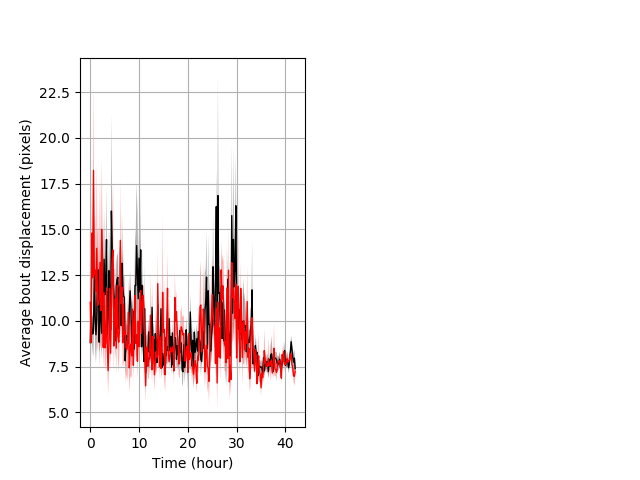
Behavior Data Description 4
Frequency of movement entire protocol Graph Pvalue = not significant Merged section Pvalue = not significant 10 het vs 10 hom ribgraph_mean_ribbonbout_dpixnumberofbouts_10min_combo.pngBehavior Data Graph 4
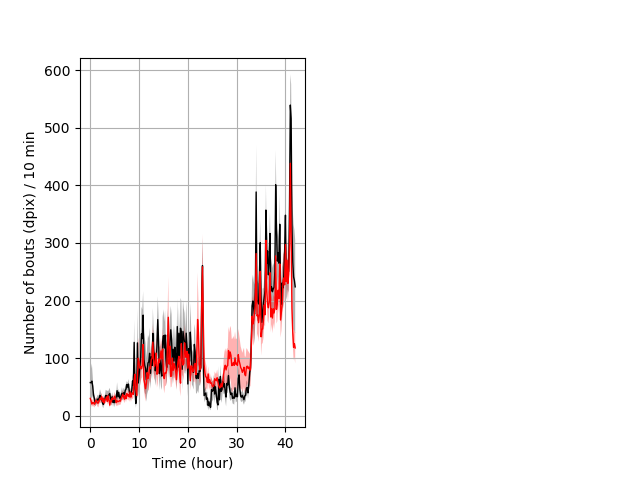
Behavior Data Description 5
Location in well entire protocol Graph Pvalue = not significant Merged section Pvalue = not significant 10 het vs 10 hom ribgraph_mean_ribbonbout_averhofrac_10min_combo.pngBehavior Data Graph 5
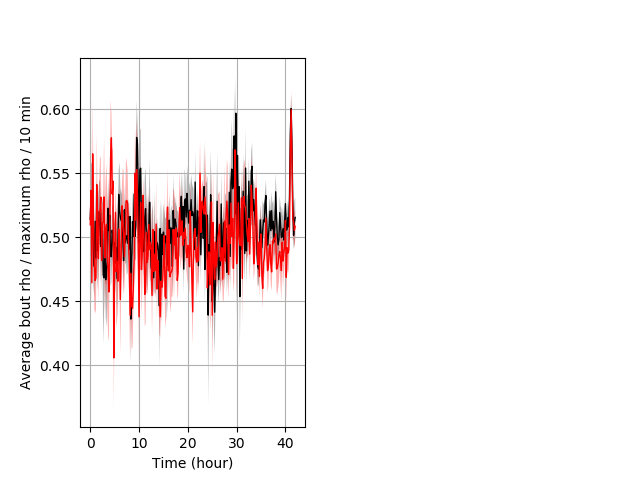
Behavior Data Description 6
NoneBehavior Data Graph 6
NoneBehavior Data Description 7
NoneBehavior Data Graph 7
NoneBehavior Data Description 8
NoneBehavior Data Graph 8
NoneBehavior Data Description 9
NoneBehavior Data Graph 9
NoneBehavior Data Description 10
NoneBehavior Data Graph 10
NoneBehavior Data Description 11
NoneBehavior Data Graph 11
NoneBehavior Data Description 12
NoneBehavior Data Graph 12
NoneBehavior Data Description 13
NoneBehavior Data Graph 13
NoneBehavior Data Description 14
NoneBehavior Data Graph 14
None