Locus Rank
3Gene
Id
10Gene Name
ARL3Duplicated
FalseMaternal
TrueGene Not Within Locus (Nearby
NoneGene Description
ADP-ribosylation factor-like 3 is a member of the ADP-ribosylation factor family of GTP-binding proteins. ARL3 binds guanine nucleotides but lacks ADP-ribosylation factor activity. Has to do with release of cargo from dynein/dynactin.Mouse Phenotype
Mice homozygous for a gene trapped allele are born at sub-Mendelian ratios, are small and sickly, fail to thrive, and die by 3 weeks of age exhibiting photoreceptor degeneration and abnormal epithelial cell proliferation and cyst formation in the kidney, liver, and pancreatic tubule structures.Additional Image
Allen Brain
Allen Regions
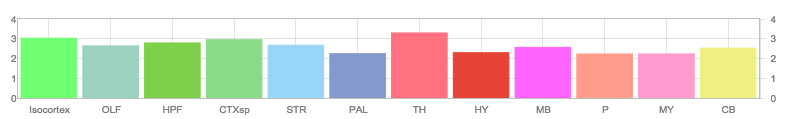
Zfin In Situ
Zfin Brain Areas
Published Zebrafish Pehnotype
Not In Allen Brain
FalseZFIN Link
NoneAllen Link
NoneMore Additional Images
Papers
Locus Rank
Locus Rank
3Genes in Region
ARL3, AS3MT, C10orf32, CNNM2, CYP17A1, INA, NT5C2, PCGF6, PDCD11, SFXN2, TAF5, TRIM8, USMG5, WBP1LAssociated Snps
rs11191419 (shown), chr10_104957618_I, rs7907645, rs55833108GWAS Region (hg19)
chr10:104423800-105165583Genes Skipping
AS3MT, NT5C2, PCGF6, PDCD11, SFXN2, TAF5, TRIM8, USMG5, WBP1LRicopili Plot Surrounding Area
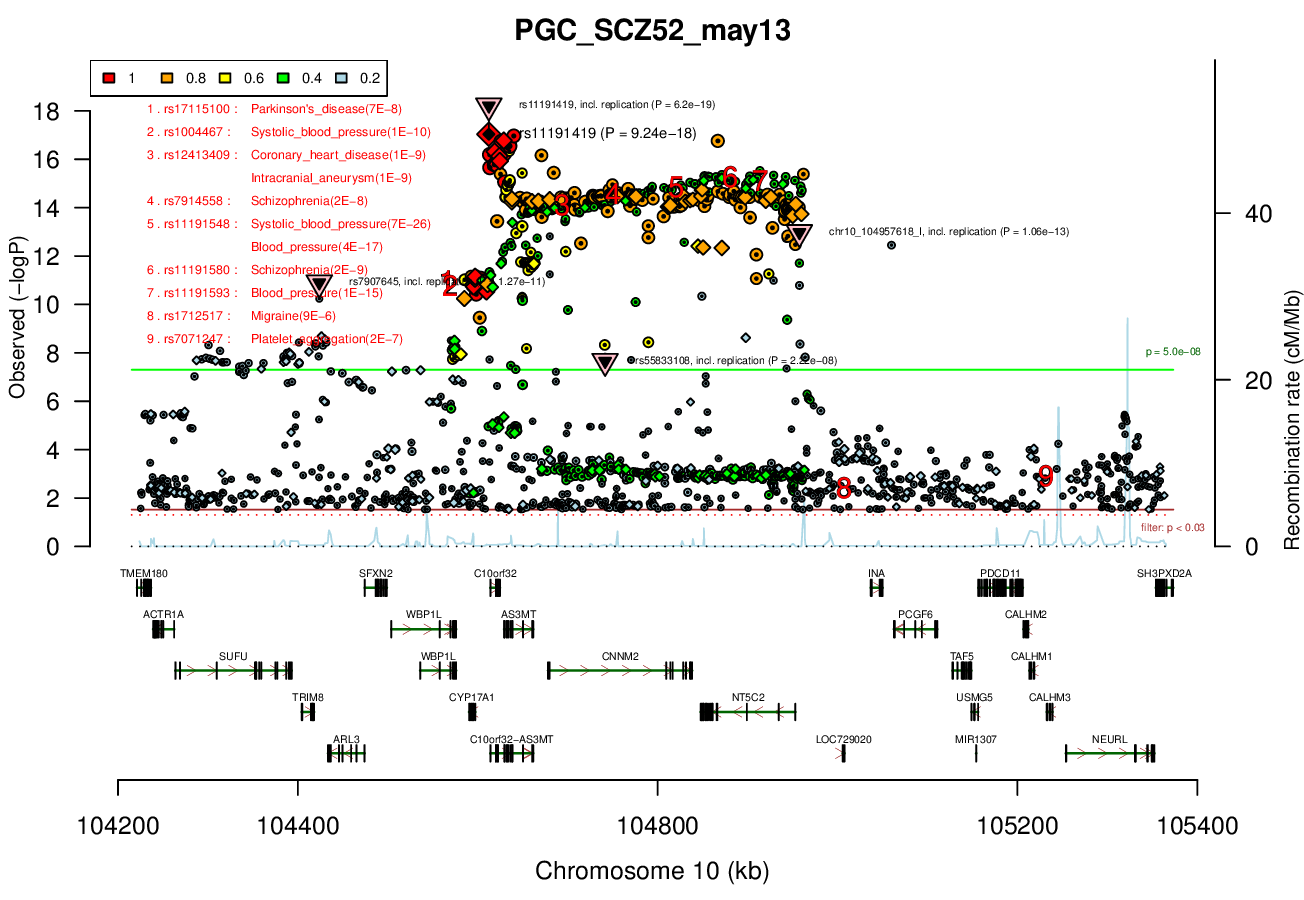
Distance On Each Side Of Locus In Above Plot
0.2 MBRicopili Plot
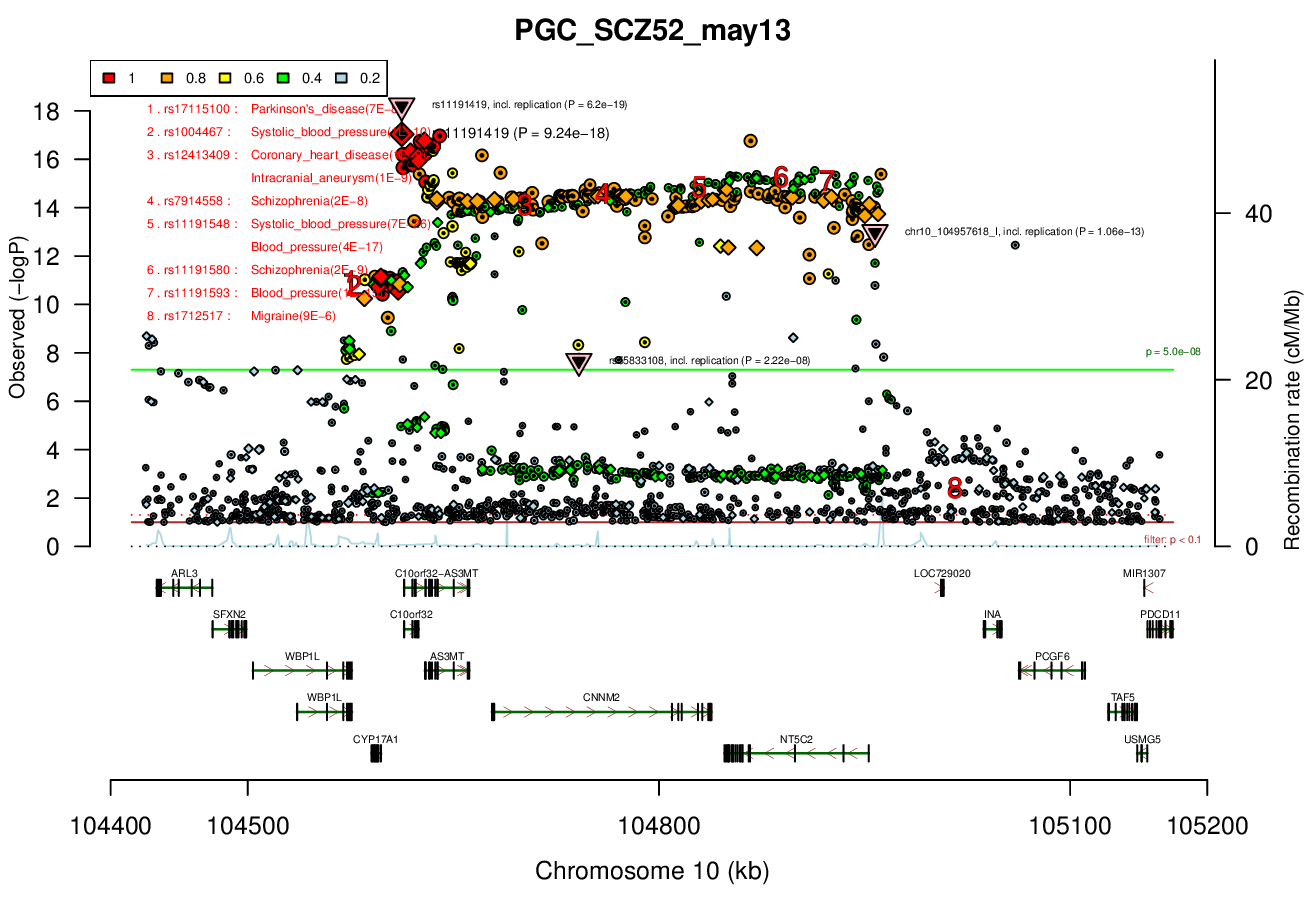
Gwas And Surrounding Region Pic
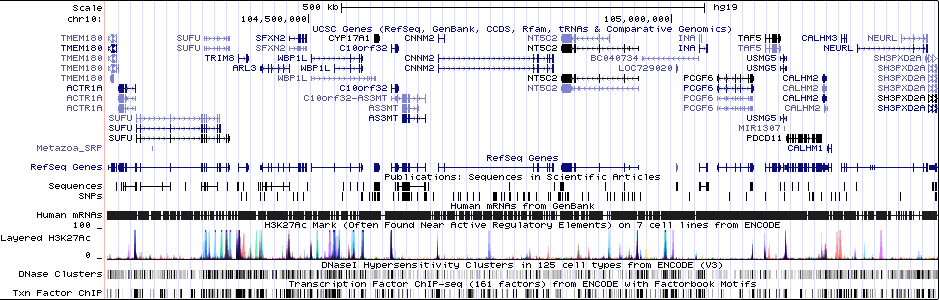
Gwas Region Pic
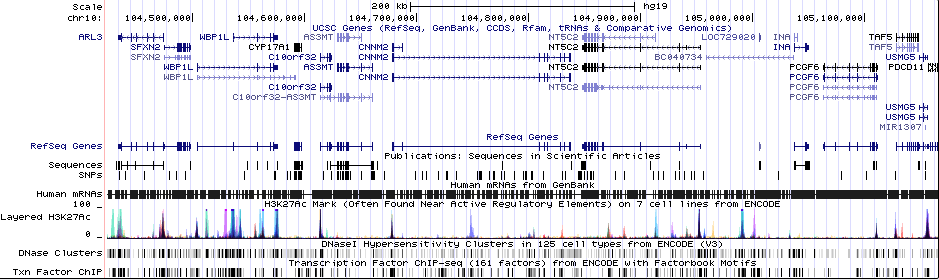
Sequence 1
Ensembl Gene Name
arl3Harvard Allele
a233Mutation Area - WT DNA Sequence
tatgacgtcatgttgcacgcattgcaaaaactcagatcactgtacattctgcgccttttagGGTTTGTTGTCCATTTTACGGAAACTGAAGAGCACCCCAGATCAGGAGGTGAGGATATTGCTTCTAGGACTGGACAATGGGGGAAAGACGACCTTACTGAAACAGCTAGCATCTGAAGACATCACTCATATCACTCCAACACAGgtacagacacggttactgtaactactgcattcacatctccagataaagctaattaMutation Area - Mutant DNA Sequence
tatgacgtcatgttgcacgcattgcaaaaactcagatcactgtacattctgcgccttttagGGTTTGTTGTCCATTTTACGGAAACTGAAGAGCACCCCAGGAGGTGAGGATATTGCTTCTAGGACTGGACAATGGGGGAAAGACGACCTTACTGAAACAGCTAGCATCTGAAGACATCACTCATATCACTCCAACACAGgtacagacacggttactgtaactactgcattcacatctccagataaagctaattaGuide RNA Target Sites
tgtacattctgcgccttttagGG
GAAGAGCACCCCAGATCAGGAGG
GCTTCTAGGACTGGACAATGGGG
CACTCATATCACTCCAACACAGg
Genotyping Primers
f, ctgcgccttttagGGTTTGTTGTCC
r, CTGTTTCAGTAAGGTCGTCTTTCCCC
Alignment of WT and Mutant Sequences w/ gRNA Targets (cyan) and Genotyping Primers (pink) Shown on WT Sequence
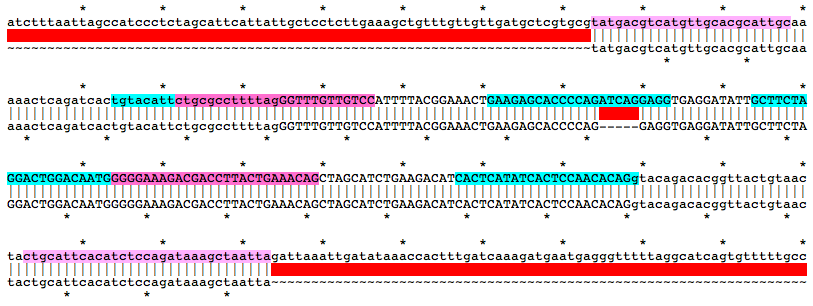
Allele
5bpDWT Genotyping Size
118WT Protein Sequence
MGLLSILRKLKSTPDQEVRILLLGLDNGGKTTLLKQLASEDITHITPTQGFNIKSVQSQG
FKLNVWDIGGQRKIRPYWRNYFENTDVLIYVIDSADRKRFEETGQELAELLDEEKLSGVP
VLVFANKQDLLTAAPASEIAEGLNLHTIRDRVWQIQSCSALTGEGVQDGMNWVCKSVNAK
RK-
Mutant Protein Sequence
MGLLSILRKLKSTPGGEDIASRTGQWGKDDLTETASI-Mutant Genotyping Size
113Ribosome Profiling Development
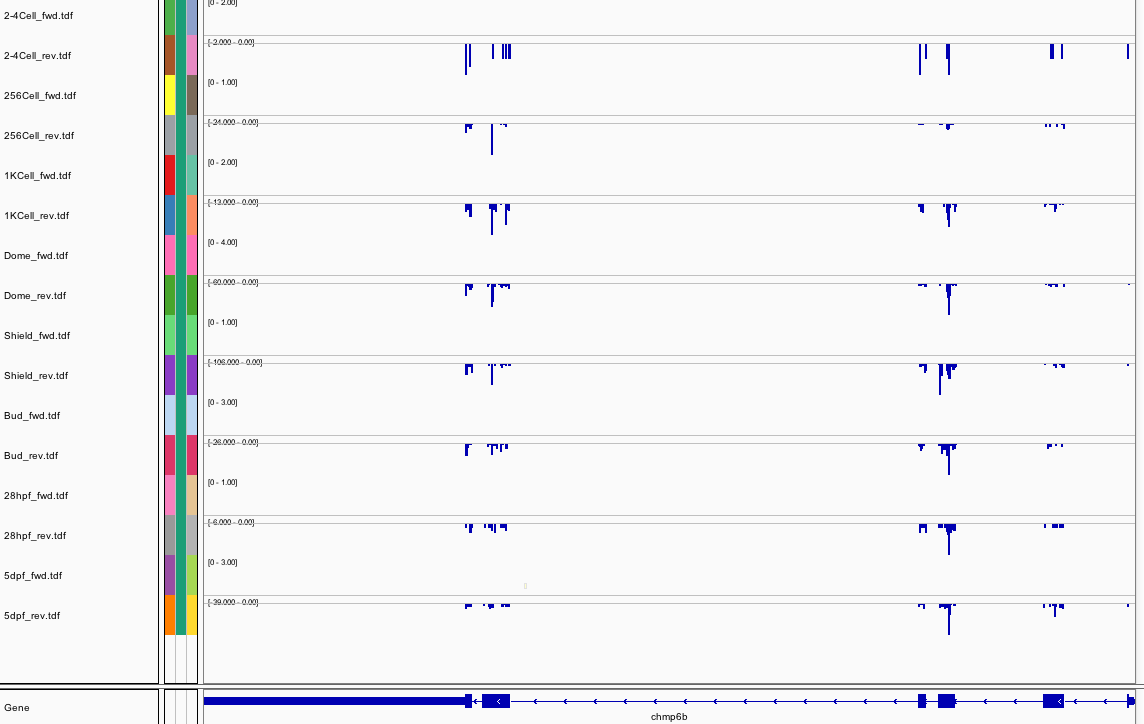
Transcript Plot
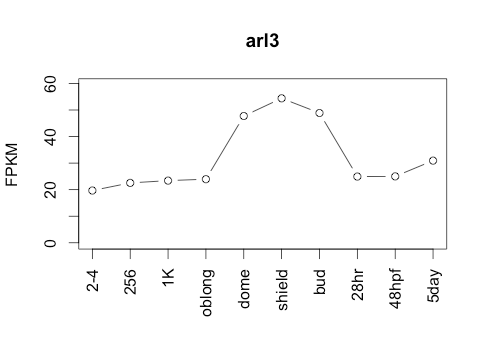
Behavior Data
Behavior Data Summary
NoneBehavior Data Description 1
Dark flash block 1 start Merged section Pvalue = not significant 30 het vs 18 hom ribgraph_mean_ribbon_fullboutdata_dpix_a0darkflash103.pngBehavior Data Graph 1
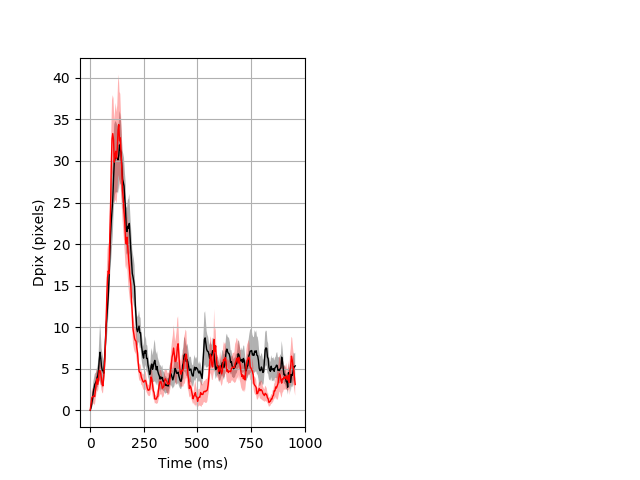
Behavior Data Description 2
Day all prepulse tap Merged section Pvalue = 0.0167 30 het vs 18 hom ribgraph_mean_ribbon_fullboutdata_dpix_dayprepulseinhibition100d.pngBehavior Data Graph 2
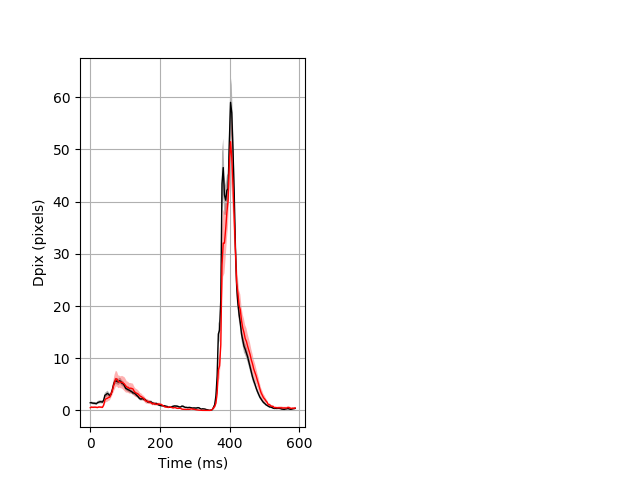
Behavior Data Description 3
Features of movement entire protocol Graph Pvalue = 0.00684 Merged section Pvalue = 0.0196 30 het vs 18 hom ribgraph_mean_ribbonbout_aveboutdisp_10min_combo.pngBehavior Data Graph 3
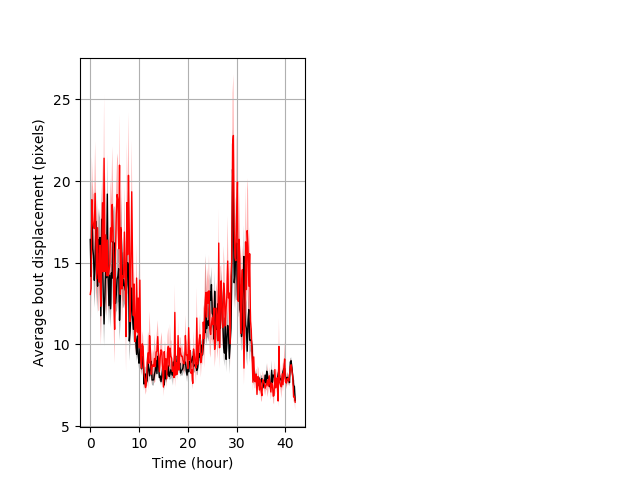
Behavior Data Description 4
Frequency of movement entire protocol Graph Pvalue = 0.000561 Merged section Pvalue = 0.00403 30 het vs 18 hom ribgraph_mean_ribbonbout_dpixnumberofbouts_10min_combo.pngBehavior Data Graph 4
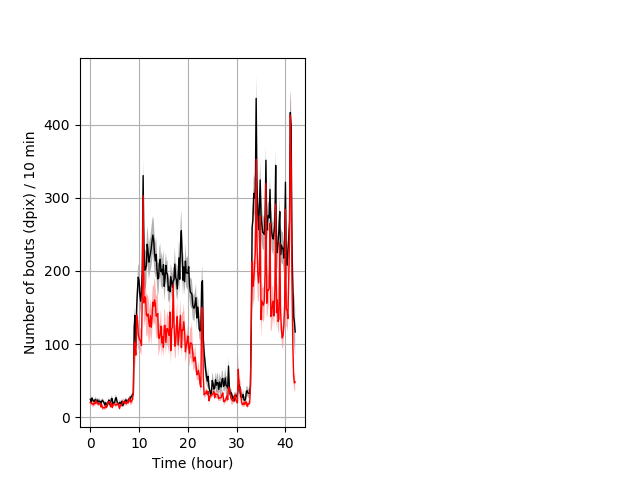
Behavior Data Description 5
Location in well entire protocol Graph Pvalue = not significant Merged section Pvalue = not significant 30 het vs 18 hom ribgraph_mean_ribbonbout_averhofrac_10min_combo.pngBehavior Data Graph 5
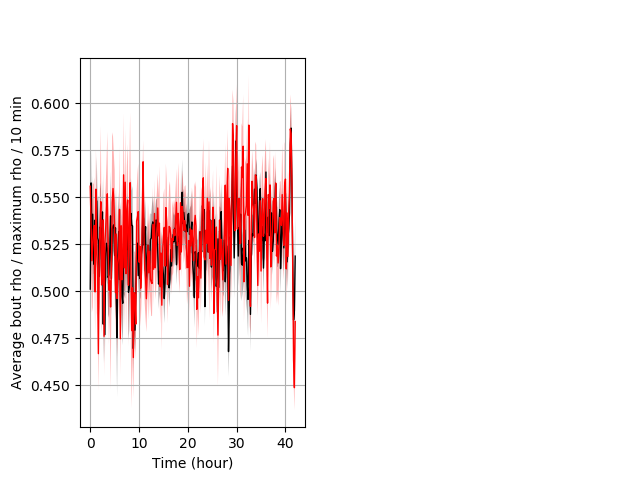
Behavior Data Description 6
NoneBehavior Data Graph 6
NoneBehavior Data Description 7
NoneBehavior Data Graph 7
NoneBehavior Data Description 8
NoneBehavior Data Graph 8
NoneBehavior Data Description 9
NoneBehavior Data Graph 9
NoneBehavior Data Description 10
NoneBehavior Data Graph 10
NoneBehavior Data Description 11
NoneBehavior Data Graph 11
NoneBehavior Data Description 12
NoneBehavior Data Graph 12
NoneBehavior Data Description 13
NoneBehavior Data Graph 13
NoneBehavior Data Description 14
NoneBehavior Data Graph 14
None