Locus Rank
3Gene
Id
6Gene Name
CNNM2Duplicated
TrueMaternal
TrueGene Not Within Locus (Nearby
FalseGene Description
This gene encodes a member of the ancient conserved domain containing protein family. Members of this protein family contain a cyclin box motif and have structural similarity to the cyclins. The encoded protein may play an important role in magnesium homeostasis by mediating the epithelial transport and renal reabsorption of Mg2+. Mutations in this gene are associated with renal hypomagnesemia. Mouse Phenotype
Additional Image
Allen Brain
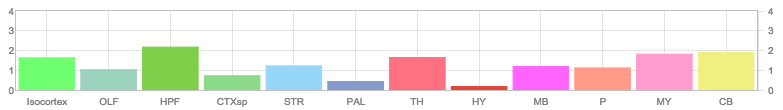
Allen Regions
Zfin In Situ
Zfin Brain Areas
cnnm2a (see in situ): Midbrain-hindbrain boundaryPublished Zebrafish Pehnotype
Not In Allen Brain
FalseZFIN Link
http://zfin.org/search?q=cnnm2&fq=category%3A%22Gene+%2F+Transcript%22&category=Gene+%2F+Transcript&fq=type%3A%22Gene%22Allen Link
http://mouse.brain-map.org/gene/show/60853More Additional Images
Papers
Locus Rank
Locus Rank
3Genes in Region
ARL3, AS3MT, C10orf32, CNNM2, CYP17A1, INA, NT5C2, PCGF6, PDCD11, SFXN2, TAF5, TRIM8, USMG5, WBP1LAssociated Snps
rs11191419 (shown), chr10_104957618_I, rs7907645, rs55833108GWAS Region (hg19)
chr10:104423800-105165583Genes Skipping
AS3MT, NT5C2, PCGF6, PDCD11, SFXN2, TAF5, TRIM8, USMG5, WBP1LRicopili Plot Surrounding Area
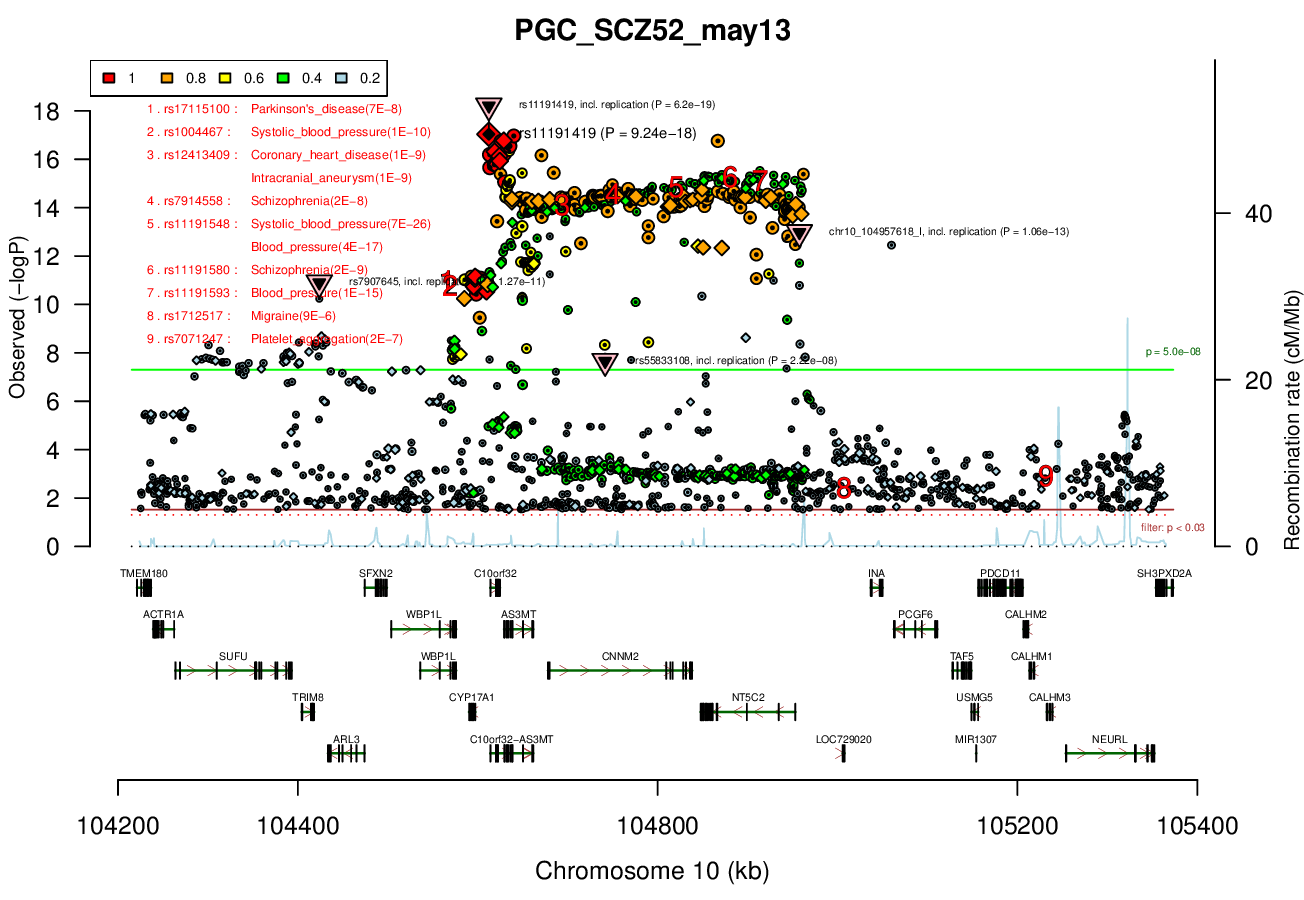
Distance On Each Side Of Locus In Above Plot
0.2 MBRicopili Plot
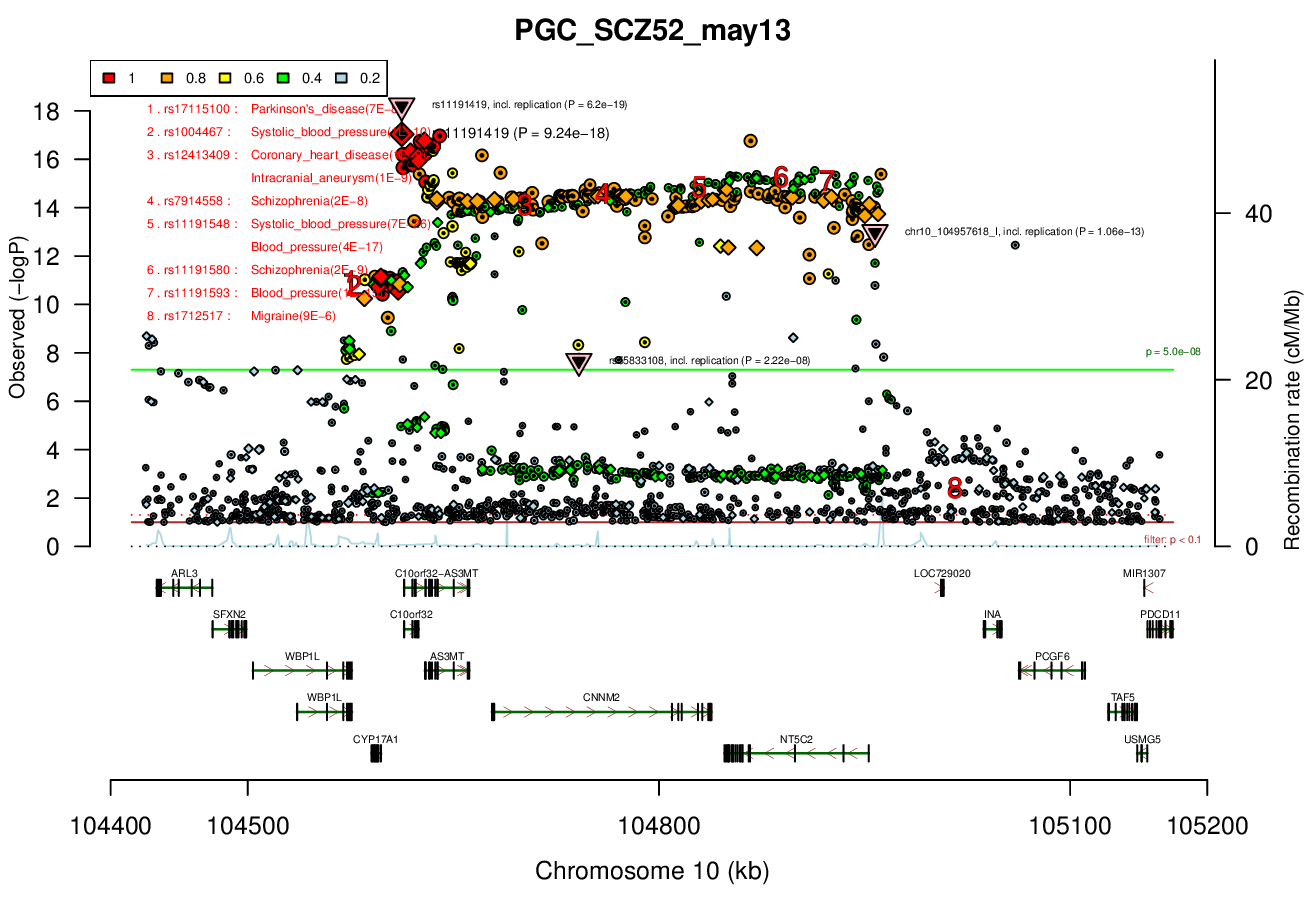
Gwas And Surrounding Region Pic
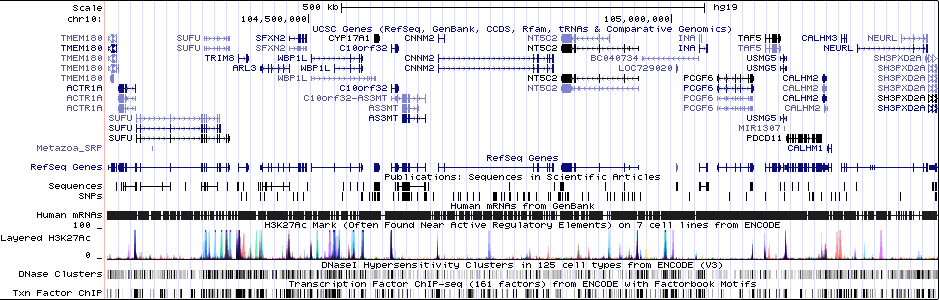
Gwas Region Pic
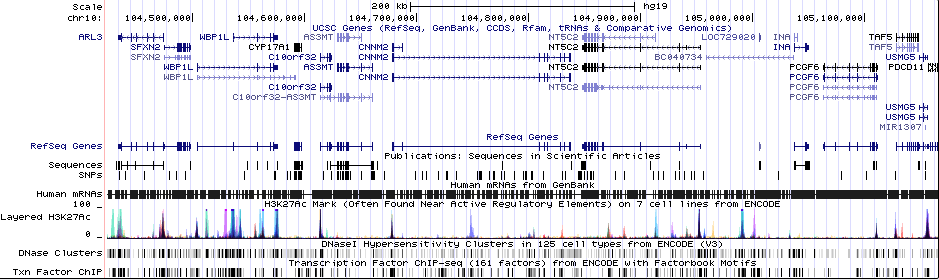
Sequence 1
Ensembl Gene Name
cnnm2aHarvard Allele
a357Mutation Area - WT DNA Sequence
CTAAACCGGGCTGGATCATTGACTGTTATATTCCTAATTGTCAGCAGTTGTCTGATCAAAAGCATTGATGGTAAAGACCCGTCAGAAGAGACTGTCATCATTGGCCTCCGCCTCGAAGACACCGACGAGGTGTCTTTTATGGACAGAGGCTTTTTGCGCGTCAGCGAGCGATCTCGTGTCAGATTAAGGGTCTATGGGCAAAACATCAACAACGAGACGTGGTCCAAACTCGCCTTCACGGAACACCGCAGCAACGCACCTGCGGAGAGCCTGAGCCCCGGAGACGCACCTGCCTCGCATCCCTGCGGCATCCGATCCTCCGATATCATCATCCTTCCCACCATGGAGGTAAACAGAAAGACTTCTGGGATTATCGAGCTCGAGGTGAAGCCTCTGCGMutation Area - Mutant DNA Sequence
CTAAACCGGGCTGGATCATTGACTGTTATATTCTTAATTGTCAGCAGTTGTCTGATCAAAAGAAGAGACTGTCATCATTGGCCTCGAAGACACCGACGAGGTGTCTTTTATGGACAGAGGCTTTTTGCGCGTCAGCGAGCGATCTCGTGTCAGATTAAGGGTCTATGGGCAAAACATCAACAACGAGACGTGGTCCAAACTCGCCTTCACGGAACACCGCAGCAACGCACCTGCGGAGAGCCTGAGCCCCGGAGACGCACCTGCCTCGCATCCCTGCGGCATCCGATCCTCCGATATCATCATCCTTCCCACCATGGAGGTAAACAGAAAGACTTCTGGGATTATCGAGCTCGAGGTGAAGCCTCTGCGGuide RNA Target Sites
TGATGACAGTCTCTTCTGACGGG
CGTCGGTGTCTTCGAGGCGGAGG
GCGAGGCAGGTGCGTCTCCGGGG
TGTTTACCTCCATGGTGGGAAGG
Genotyping Primers
f, GCTAAACCGGGCTGGATCATTGACTGT
r, CGCAGAGGCTTCACCTCGAGCT
Alignment of WT and Mutant Sequences w/ gRNA Targets (cyan) and Genotyping Primers (pink) Shown on WT Sequence
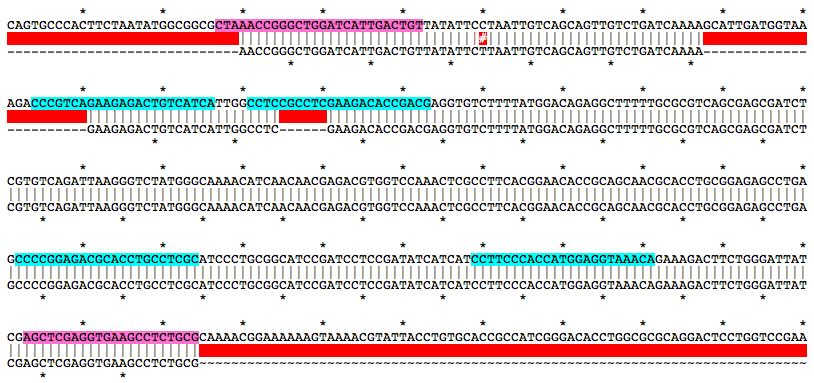
Allele
23bpD,6bpDWT Genotyping Size
398WT Protein Sequence
MAEQWTAVPTSNMAALNRAGSLTVIFLIVSSCLIKSIDGKDPSEETVIIGLRLEDTDEVS
FMDRGFLRVSERSRVRLRVYGQNINNETWSKLAFTEHRSNAPAESLSPGDAPASHPCGIR
SSDIIILPTMEVNRKTSGIIELEVKPLRKTEKSKTYYLCTAIGTPGAQDSWSEHTWVYHE
GHDTKVIVVEEKKFLLPFWLQVIFIAMLLCLSGMFSGLNLGLMALDPMELRIVQNCGTEK
EKHYAKAIEPVRSQGNYLLCSLLLGNVLVNTTLTILLDDIAGSGLIAVVVSTIGIVIFGE
IVPQAICSRHGLAVGANTIFLTKFFMILTFPASYPVSKLLDHVLGQEIGTVYNREKLLEM
LRVTDPYNDLVKEELNIIQGALELRTKTVEDVMTPLRDCFMISGDAILDFATMSEIMESG
YTRIPVYEGERCHIVDLLFVKDLAFVDPDDCTPLKTITKFYSHPLHFVFNDTKLDAMLEE
FKKGKSHLAIVQRVNNEGEGDPFYEVLGIVTLEDVIEEIIKSEILDETDLYTDNKTKKKI
AHREKKQDFSAFKPTENELRVKISPQLLLATLRFLATEVESFSATQMSEKILLRLLKHPN
VIQELKYDDKDKKMSEHYLFHRNKPVDYFILILQGKVEVEAGKEGMKFEAGAFSYYGVLS
LTSSPVPLSLSRTFVVSRAESLAGSPENKSPPRPFGLNHSDSLNRSDRMEAVTPTLGSSN
NQLNSFLQVYVPDYSVRAVSDLLYIKVTRQQYQNALMASRMDKTPQSTDSEFTKIELTFT
EHQDGPLEESATLLTTNSHKPNHTAHNDGVV-
Mutant Protein Sequence
MAEQWTAVPTSNMAALNRAGSLTVIFLIVSSCLIKRRDCHHWPRRHRRGVFYGQRLFARQRA
ISCQIKGLWAKHQQRDVVQTRLHGTPQQRTCGEPEPRRRTCLASLRHPILRYHHPSHHGGK
QKDFWDYRARGEASAQNGKK-
In Situ Probe Sequence
ATTGCCATGTTGTTGTGCTTGTCTGGGATGTTCAGCGGGCTGAATCTGGGTCTGATGGCGCTGGATCCCATGGAGCTCCGGATTGTGCAAAACTGCGGGACCGAGAAGGAGAAGCATTACGCGAAGGCGATCGAGCCTGTCCGAAGCCAAGGAAACTACCTGCTGTGCTCGTTGCTGCTGGGAAATGTCCTGGTCAACACTACTTTGACTATACTTCTCGATGACATCGCAGGGTCAGGGCTGATCGCTGTGGTGGTGTCCACTATAGGGATAGTGATATTTGGGGAGATCGTGCCTCAGGCCATCTGCTCCAGGCACGGACTCGCTGTAGGCGCGAATACAATATTCTTGACTAAATTTTTTATGATCCTCACTTTCCCAGCATCCTATCCGGTCAGCAAACTCCTGGACCACGTGCTGGGGCAGGAGATTGGAACTGTGTACAACAGGGAGAAGCTGCTGGAGATGCTGAGGGTGACCGATCCATATAATGACCTGGTCAAAGAGGAGCTCAACATCATCCAGGGCGCTCTGGAGCTCAGGACTAAAACCGTGGAGGATGTCATGACACCTCTGCGGGACTGCTTCATGATTTCTGGTGATGCCATACTGGACTTCGCCACCATGTCTGAGATCATGGAGAGTGGCTACACTCGGATACCTGTGTATGAGGGTGAGAGATGTCACATTGTGGACCTGCTGTTTGTCAAGGACTTGGCCTTTGTGGATCCAGATGACTGCACGCCTCTGAAAACCATCACCAAGTTCTACAGCCACCCTCTGCACTTTGTCTTCAACGATACAAAGCTGGATGCCATGCTGGAGGAGTTTAAAAAAGGTAAATCCCATCTGGCCATTGTCCAAAGAGTGAATAATGAAGGAGAGGGAGACCCGTTCTATGAAGMutant Genotyping Size
369Ribosome Profiling Development
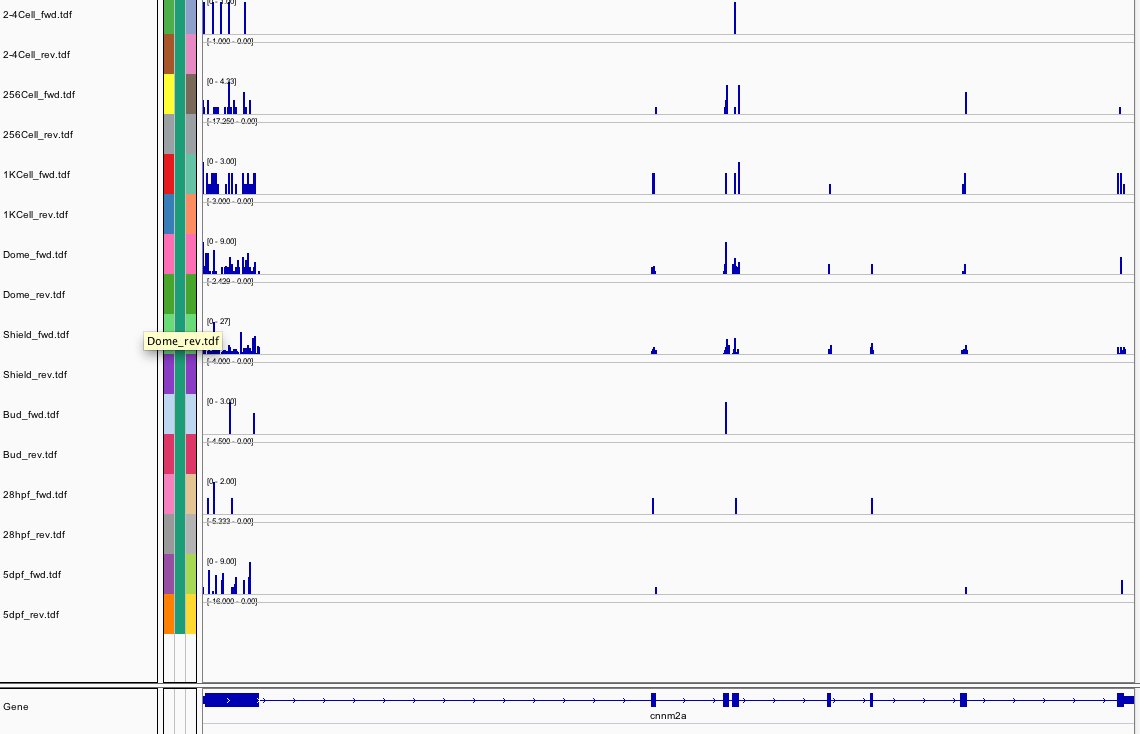
Transcript Plot
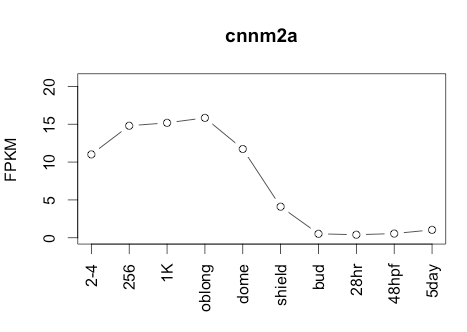
Sequence 2
Ensembl Gene Name
cnnm2bHarvard Allele
a258Mutation Area - WT DNA Sequence
CAGAACCGTCGGTTCCAGTACCCCCTCGAAAAATGGCGTCACTCAGCCGGGTCCGAGCCTTTACTCTGATTTTCTTAACTGCAAGCAGTTGTTTAATTACTCCCACAAGTGGCAAGGAAGGAGGGGGAGAGGAGACGGTCATCATCGGGCTGCGGCTGGAGGACACGGACGACATTTCCTTCATGGACAGGGGCTTTTTGCGGGTCAGCGAGAGGTCTCGGGTGAGATTGAGGGTGTACGGGCAAAACATCAACAACGAGACTTGGTCCAAAATCGCATTTACGGAGCACGAGAGGAGCCGAGTGATCGGCAACATCGACAGCTCTTCGGGGGACAACCAGAGTCAGGAGGACACGTCGGGCTTGCACCCCTGCGGCATCAGAACTTCGGATATTCTGATCTTGCCCAACATCGTTTTAAACCGCAAGACGTCTGGCACCGTGGAGATAGAGGTCAAACCTCTGCGAAAGACTGAGCGGAGCAAAGCGTACTACCTGTGCATCGCCACCTCCACACCTGCTGTGGCCGGAGCGCACGACCCGTGGACCGAGAACACGTGGATCTATCACGATGGAGAGGACACCAAAGTGATCGTGGTGGAGGAGAAAAAGTTCCTGCTGCCCTTCTGGCTCCAGGTCATCTTCATTTCTATGCTGCTCTGCCTCTCGGGCATGTTCAGCGGCTTGAACTTGGGGCTGATGGCGCTGGACCCCATGGAGMutation Area - Mutant DNA Sequence
CAGAACCGTCGGTTCCAGTACCCCCTCGAAAAATGGCGTCACTCAGCCGGGTCCGAGCCTTTACTCTGATTTTCTTAACTGCAAGCAGTTGTTTAATTACTCCCACAAGTGGCAAGGAAGCATCTTCAGGACCTGGAGCATCTTCATTTCTATGCTGCTCTGCCTCTCGGGCATGTTCAGCGGCTTGAACTTGGGGCTGATGGCGCTGGACCCCATGGAGGuide RNA Target Sites
TGGCAAGGAAGGAGGGGGAGAGG
GCAACATCGACAGCTCTTCGGGG
CCGCAAGACGTCTGGCACCGTGG
GAAGATGACCTGGAGCCAGAAGG
Genotyping Primers
f, CAGAACCGTCGGTTCCAGTACCCC
r, CTCCATGGGGTCCAGCGCC
wt_f, TAGAGGTCAAACCTCTGCGAAAGACTG
PCR can be run with mix of all three primers.
Alignment of WT and Mutant Sequences w/ gRNA Targets (cyan) and Genotyping Primers (pink) Shown on WT Sequence
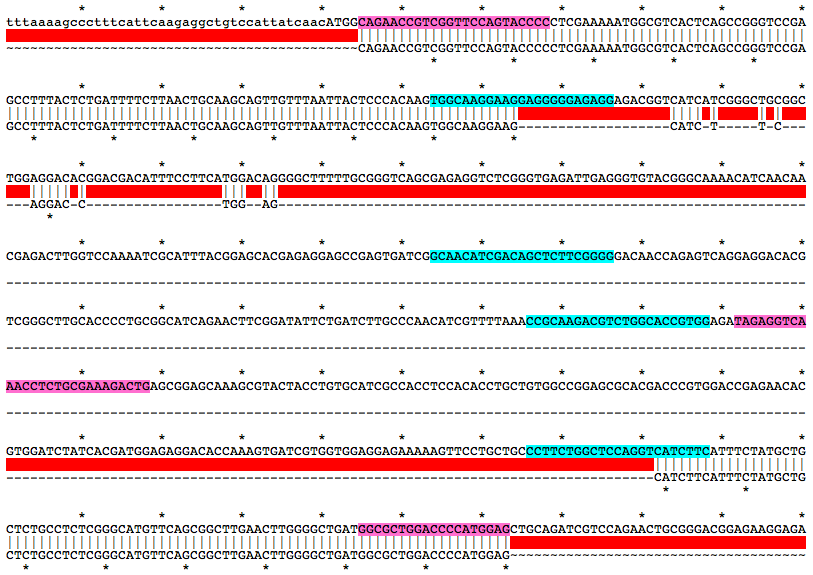
Allele
499bpDWT Genotyping Size
272WT Protein Sequence
MAEPSVPVPPRKMASLSRVRAFTLIFLTASSCLITPTSGKEGGGEETVIIGLRLEDTDDI
SFMDRGFLRVSERSRVRLRVYGQNINNETWSKIAFTEHERSRVIGNIDSSSGDNQSQEDT
SGLHPCGIRTSDILILPNIVLNRKTSGTVEIEVKPLRKTERSKAYYLCIATSTPAVAGAH
DPWTENTWIYHDGEDTKVIVVEEKKFLLPFWLQVIFISMLLCLSGMFSGLNLGLMALDPM
ELQIVQNCGTEKEKNYARKIEPVRSQGNYLLCSLLLGNVLVNTTLTILLDDIAGSGLIAV
VMSTIGIVIFGEIVPQAICSRHGLAVGANTIFLTKFFMLLTFPASYPVSKLLDYLLGQEI
GTVYNREKLLEMLRVTDPYNDLVKEELNIIQGALELRTKTVEDVMTPLRDCFMITGDATL
DFNSMSEIMESGYTRIPVFEGDRSNIVDLLFVKDLAFVDPDDCTPLKTITKFYSHPLHFV
FNDTKLDAMLEEFKKGKSHLAIVQRVNNEGEGDPFYEVLGIVTLEDVIEEIIKSEILDET
DLYTDNKTKKKITHRERKQDFSAFKHMDNEMKVKISPQLLLATLRFLATEVEPFSPSQMS
EKILLRLLKHPNVIQEVKYNDKNKKAIEHYLFHRNKPVDYFILVLQGKVEVEAGKEGLRF
EAGAFSSYGMMALTASPENKSPPRSFGLNHSDSLNRSERIEVITPSLGSSNNQLNAFLQI
YVPDYSVRAVSDVQYIKVTRQQYQNAVMASRIDKTPQSSESEYTKIEMALTELHDGLTDE
TTTLFNQTQNCVVHNKSNHTGHSEGPI-
Mutant Protein Sequence
MAEPSVPVPPRKMASLSRVRAFTLIFLTASSCLITPTSGKEASSGPGASSFLCCSASRACSAA-In Situ Probe Sequence
ATGGCAGAACCGTCGGTTCCAGTACCCCCTCGAAAAATGGCGTCACTCAGCCGGGTCCGAGCCTTTACTCTGATTTTCTTAACTGCAAGCAGTTGTTTAATTACTCCCACAAGTGGCAAGGAAGGAGGGGGAGAGGAGACGGTCATCATCGGGCTGCGGCTGGAGGACACGGACGACATTTCCTTCATGGACAGGGGCTTTTTGCGGGTCAGCGAGAGGTCTCGGGTGAGATTGAGGGTGTACGGGCAAAACATCAACAACGAGACTTGGTCCAAAATCGCATTTACGGAGCACGAGAGGAGCCGAGTGATCGGCAACATCGACAGCTCTTCGGGGGACAACCAGAGTCAGGAGGACACGTCGGGCTTGCACCCCTGCGGCATCAGAACTTCGGATATTCTGATCTTGCCCAACATCGTTTTAAACCGCAAGACGTCTGGCACCGTGGAGATAGAGGTCAAACCTCTGCGAAAGACTGAGCGGAGCAAAGCGTACTACCTGTGCATCGCCACCTCCACACCTGCTGTGGCCGGAGCGCACGACCCGTGGACCGAGAACACGTGGATCTATCACGATGGAGAGGACACCAAAGTGATCGTGGTGGAGGAGAAAAAGTTCCTGCTGCCCTTCTGGCTCCAGGTCATCTTCATTTCTATGCTGCTCTGCCTCTCGGGCATGTTCAGCGGCTTGAACTTGGGGCTGATGGCGCTGGACCCCMutant Genotyping Size
220Ribosome Profiling Development
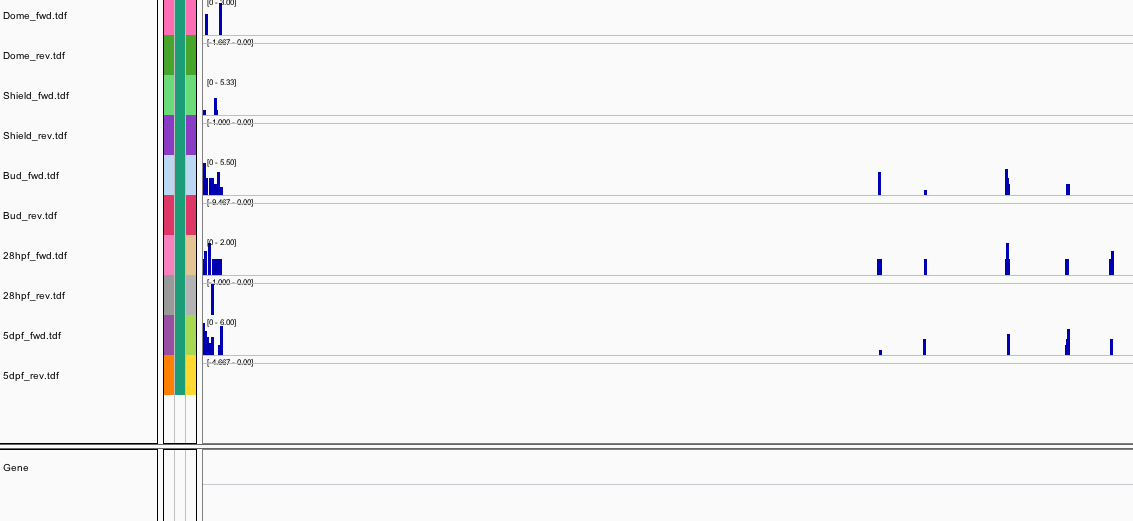
Behavior Data
Behavior Data Summary
NoneBehavior Data Description 1
Dark flash block 1 start Merged section Pvalue = 0.0184 16 hethom vs 12 homhom ribgraph_mean_ribbon_fullboutdata_dpix_a0darkflash103.pngBehavior Data Graph 1
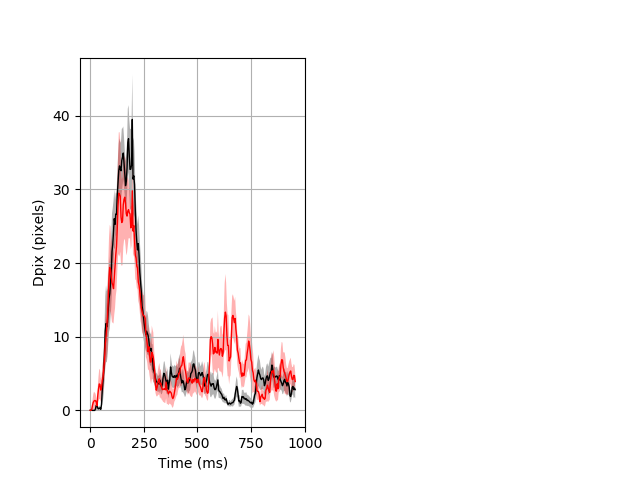
Behavior Data Description 2
Day all prepulse tap Merged section Pvalue = 0.0129 16 hethom vs 12 homhom ribgraph_mean_ribbon_fullboutdata_dpix_dayprepulseinhibition100d.pngBehavior Data Graph 2
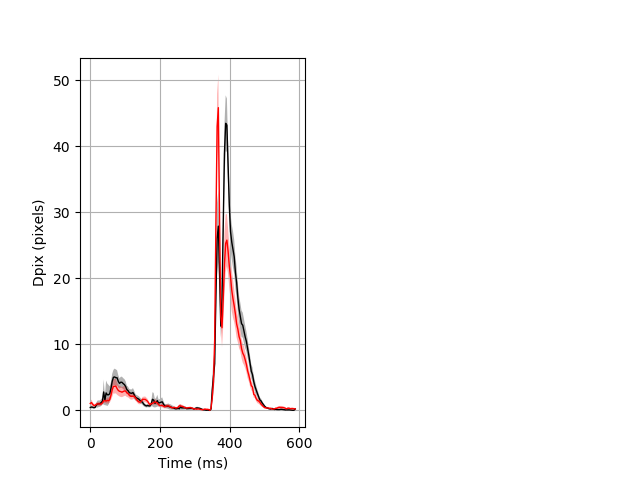
Behavior Data Description 3
Features of movement entire protocol Graph Pvalue = 0.0208 Merged section Pvalue = 0.0226 might not repeat 18 hethet vs 23 homhet ribgraph_mean_ribbonbout_aveboutdisp_10min_combo.pngBehavior Data Graph 3
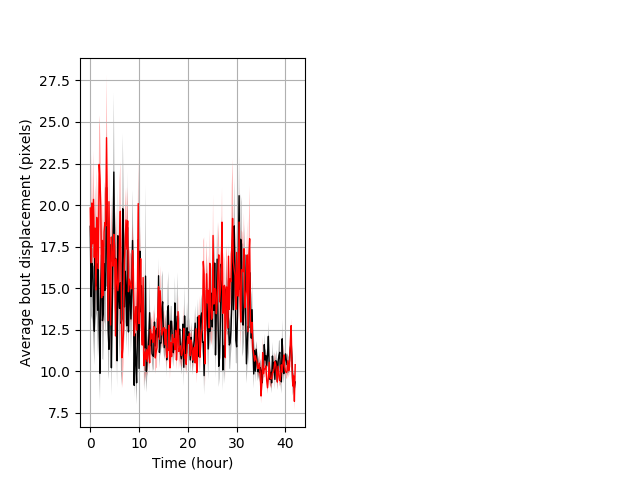
Behavior Data Description 4
Frequency of movement entire protocol Graph Pvalue = 0.0107 Merged section Pvalue = 0.0134 16 hethom vs 12 homhom ribgraph_mean_ribbonbout_dpixnumberofbouts_10min_combo.pngBehavior Data Graph 4
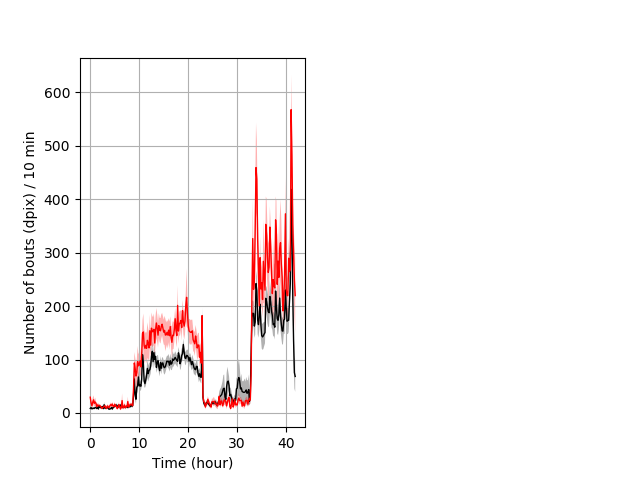
Behavior Data Description 5
Location in well entire protocol Graph Pvalue = 0.00252 Merged section Pvalue = 0.00649 18 hethet vs 23 homhom ribgraph_mean_ribbonbout_averhofrac_10min_combo.pngBehavior Data Graph 5
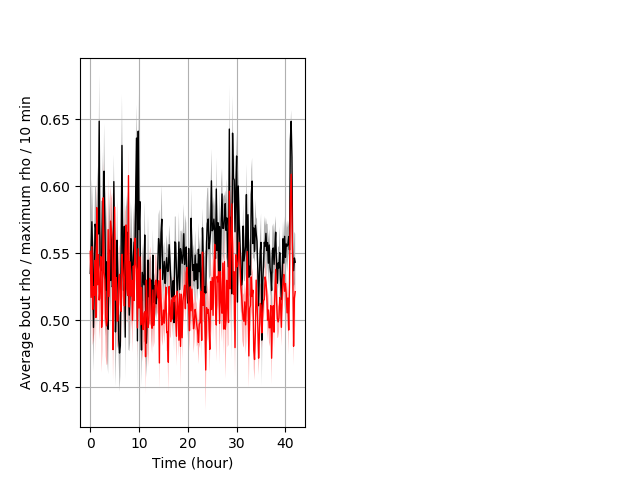
Behavior Data Description 6
Dark flash Graph Pvalue = 0.00231 18 hethet vs 23 homhom boxgraph_pd__ribgraph_mean_ribbon_fullboutdatamax_dist_ddarkflash103.pngBehavior Data Graph 6
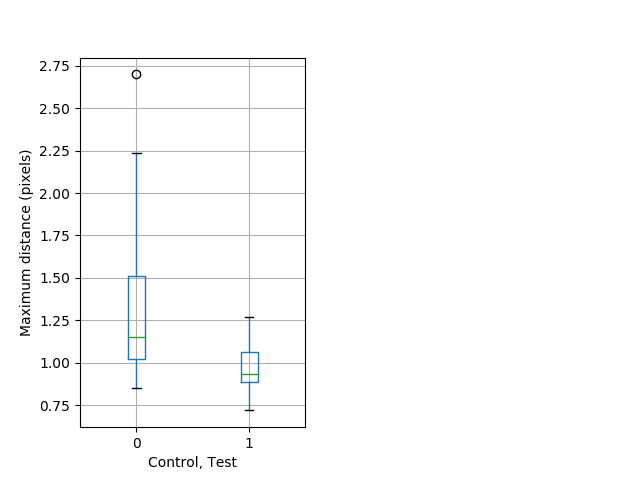
Behavior Data Description 7
Features of movement Graph Pvalue = 0.00301 17 homhet vs 12 homhom ribgraph_mean_ribbonbout_aveboutdist_min_combo.pngBehavior Data Graph 7
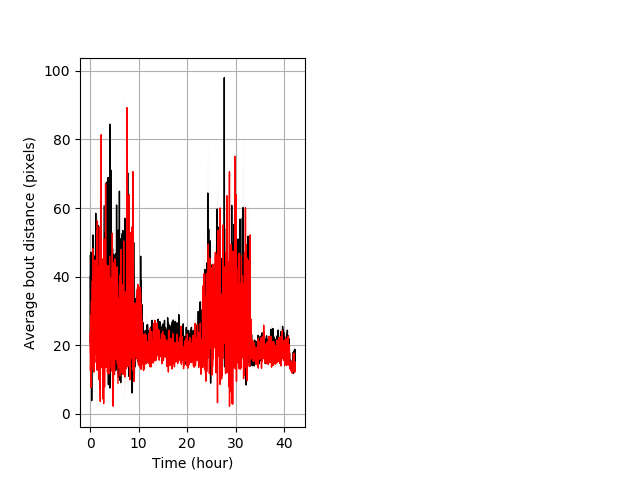
Behavior Data Description 8
Frequency of movement Graph Pvalue = 0.001 14 hethet vs 16 hethom ribgraph_mean_ribbonbout_dpixnumberofbouts_min_day2night1.pngBehavior Data Graph 8
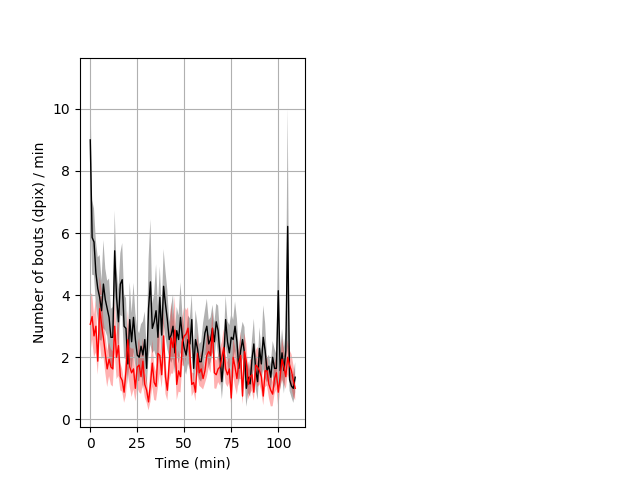
Behavior Data Description 9
Location in well Graph Pvalue = 0.00123 18 hethet vs 23 homhom ribgraph_mean_ribbonbout_averhofrac_min_combo.pngBehavior Data Graph 9
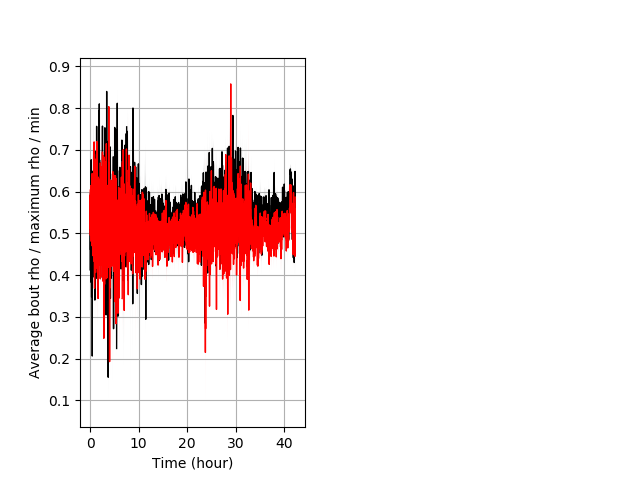
Behavior Data Description 10
PPI tap Graph Pvalue = 0.000592 16 hethom vs 12 homhom boxgraph_pd__ribgraph_mean_ribbon_latencyresponse_dpix_dayprepulseinhibition100b.pngBehavior Data Graph 10
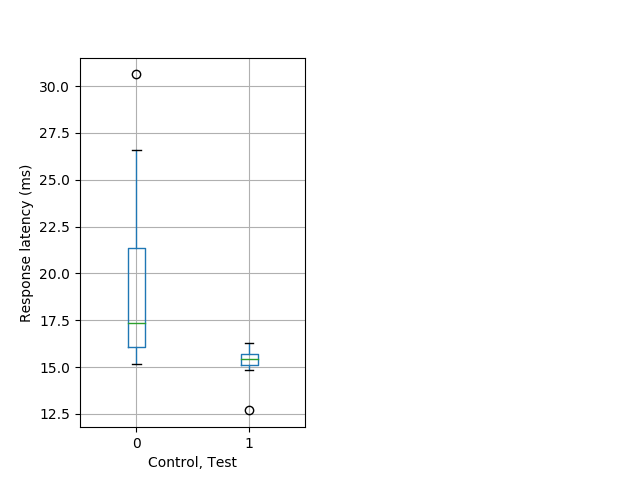
Behavior Data Description 11
Strong tap Graph Pvalue = 0.00417 14 hethet vs 17 homhet boxgraph_pd__bigmovesribgraph_mean_ribbon_peakspeed_nightprepulseinhibition102.pngBehavior Data Graph 11
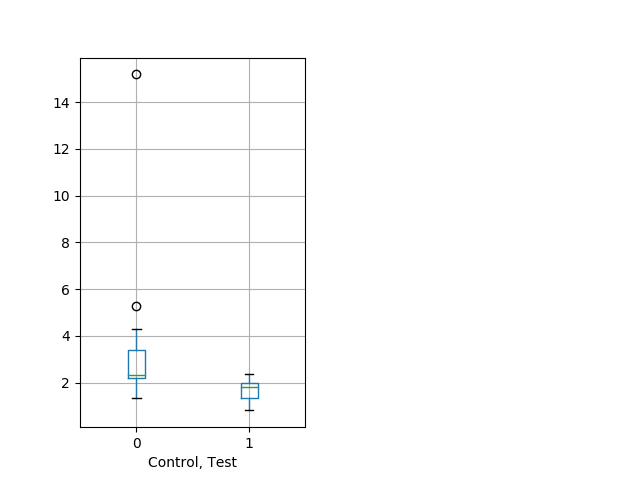
Behavior Data Description 12
Tap habituation Graph Pvalue = 0.0102 16 hethom vs 12 homhom boxgraph_pd__ribgraph_mean_ribbon_freqresponse_dpix_nighttaphab102.pngBehavior Data Graph 12
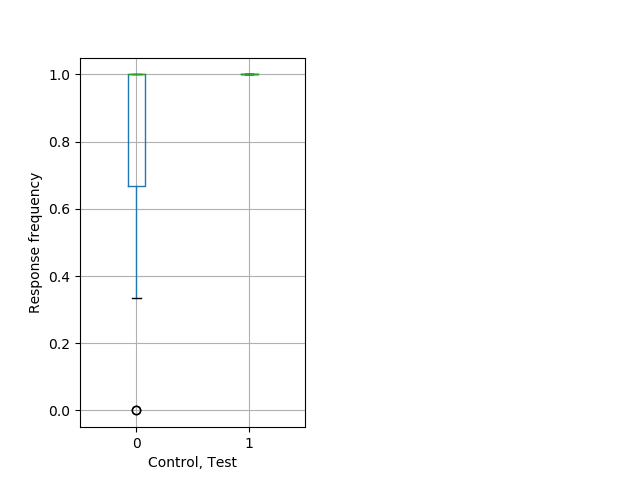
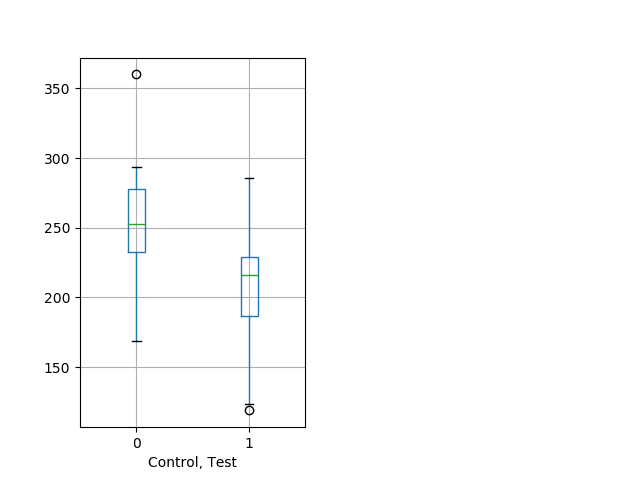
Behavior Data Description 13
Weak tap Graph Pvalue = 0.00366 17 hethet vs 17 homhom boxgraph_pd__ribgraph_mean_ribbon_peakdpix_nightprepulseinhibition100a.pngBehavior Data Graph 13
Behavior Data Description 14
NoneBehavior Data Graph 14
None