Locus Rank
3Gene
Id
122Gene Name
INADuplicated
TrueMaternal
FalseGene Not Within Locus (Nearby
FalseGene Description
Neurofilaments are type IV intermediate filament heteropolymers composed of light, medium, and heavy chains. Neurofilaments comprise the axoskeleton and they functionally maintain the neuronal caliber. They may also play a role in intracellular transport to axons and dendrites. This gene is a member of the intermediate filament family and is involved in the morphogenesis of neurons.Mouse Phenotype
Additional Image
Allen Brain
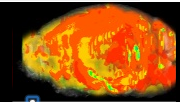
Allen Regions
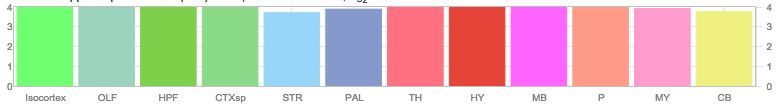
Zfin In Situ
Zfin Brain Areas
Published Zebrafish Pehnotype
Not In Allen Brain
FalseZFIN Link
NoneAllen Link
NoneMore Additional Images
Papers
http://journals.plos.org/plosone/article?id=10.1371/journal.pone.0088631Locus Rank
Locus Rank
3Genes in Region
ARL3, AS3MT, C10orf32, CNNM2, CYP17A1, INA, NT5C2, PCGF6, PDCD11, SFXN2, TAF5, TRIM8, USMG5, WBP1LAssociated Snps
rs11191419 (shown), chr10_104957618_I, rs7907645, rs55833108GWAS Region (hg19)
chr10:104423800-105165583Genes Skipping
AS3MT, NT5C2, PCGF6, PDCD11, SFXN2, TAF5, TRIM8, USMG5, WBP1LRicopili Plot Surrounding Area
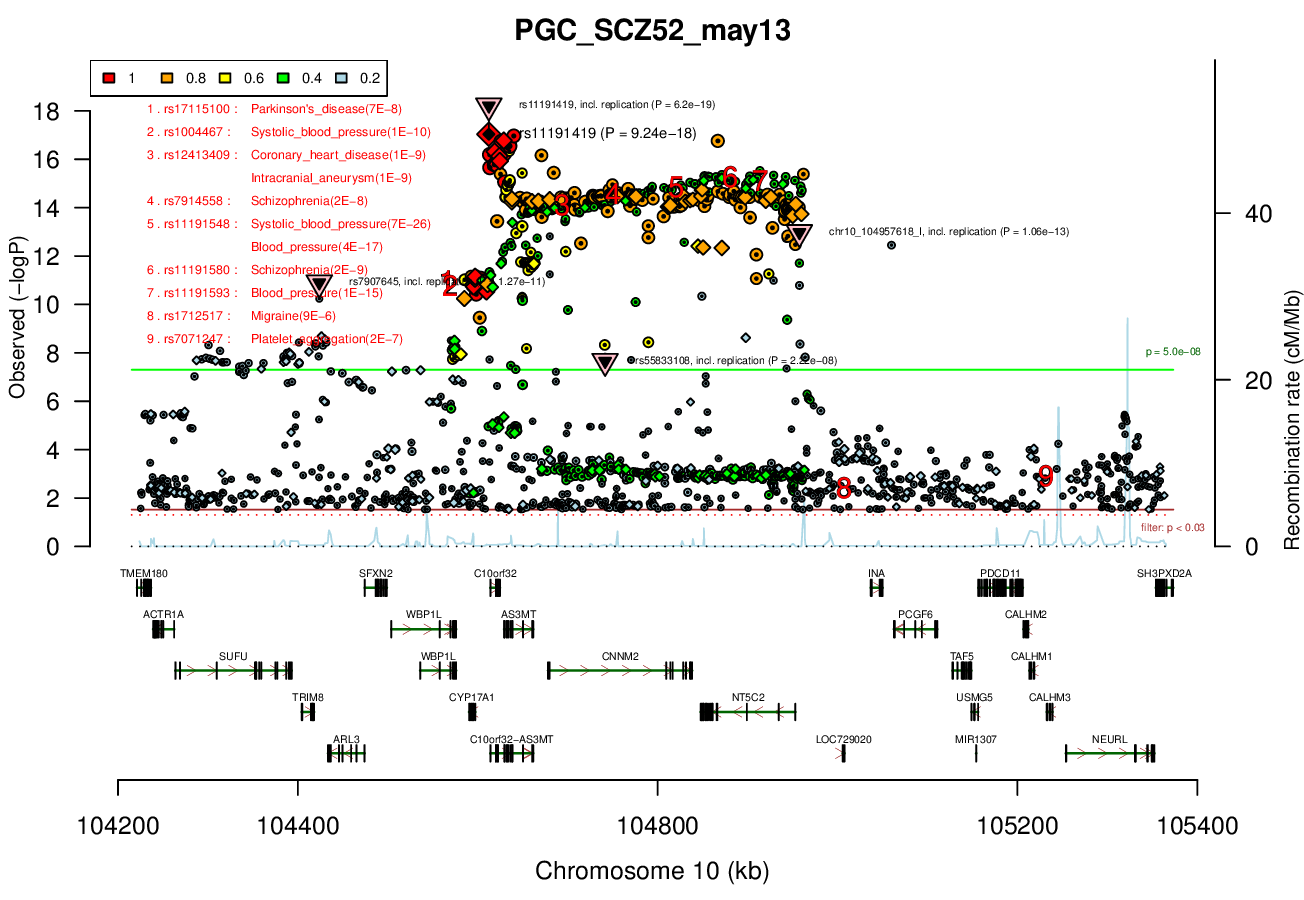
Distance On Each Side Of Locus In Above Plot
0.2 MBRicopili Plot
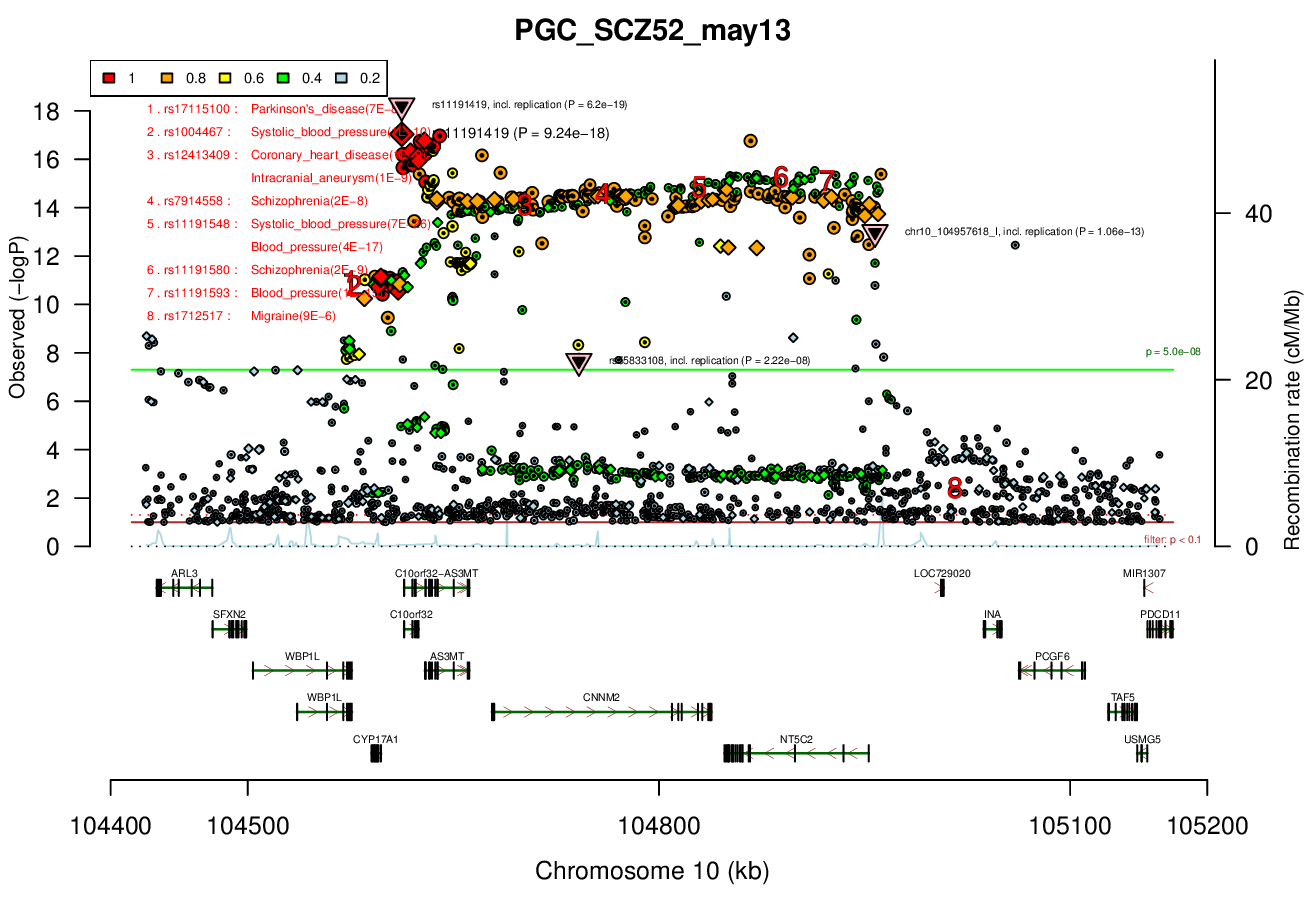
Gwas And Surrounding Region Pic
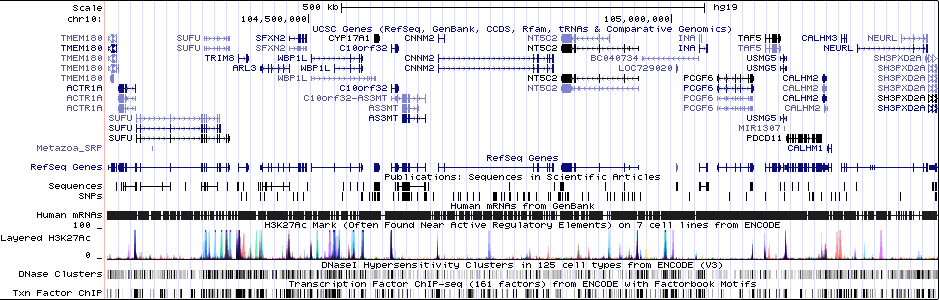
Gwas Region Pic
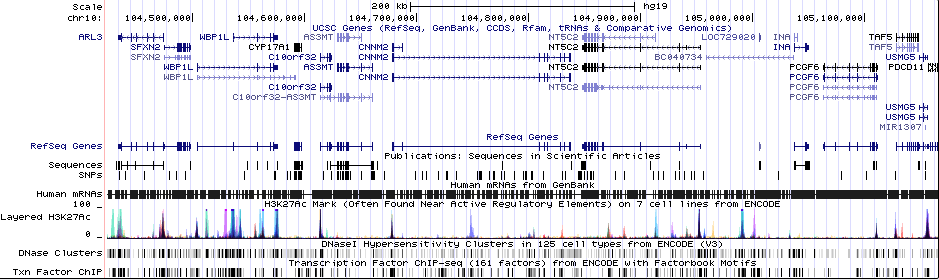
Sequence 1
Ensembl Gene Name
inaaHarvard Allele
a285Mutation Area - WT DNA Sequence
gatcttgtcttaacagtcactaccagagggaatatgacaggctgctaaatgggacagtccgccgggtatataatggcaagcacaccagttttagactcccttagttgtgacGAAGATGAGTTACGGATCAGACCACTACATGTCTTCATCCTACAGAAAATTGTTCGGGGACAACCCTCGTTTTGCCGGCTCTAGAATGAGCAATGTCTCTCCCCGGAGCTCCATGTCCTCCAGCGGCTTCAGGTCTCAGTCTGTCTCCCGCAGCAGCGCCTCTCCTGCGGGCTACTATAAAAGATCGGGCCGGTCATCTTCATTCTCGCCGGTCCATTTCGACAGCGTGGATTTCTCTCAAACCTCTGTTTTGAACAACGAGTTTAAAATCGTTCGAACCAATGAAAAGGAACAGCTGCAAGGACTCAATGACCGGTTCGCGATGTTTATCGAGAAAGTGCGCAATCTGGAGCAGCACAATAAAGTGCTGGAGACCGAACTGGTGAGTCTGCGCCAGAGGCAGAACGAGCCGTCCAGACTTGCTGAACTCTATCAGCAGGAGATCCGGGACCTGCGAGCTCAGGTAGACGAGCTGAACAACGAGAAATCGCACATTCTCATTGAACGCGACAGCATCGAGGAAGACTTGCAGAAACTTCGAGGCAAGTTTGAGGAAGAGATCCGAGCGCGCGAGGAGGCAGAGCAGACGCTCAGATCCTACAAGAAAGATGTGGATGATGCCACCATGGTCCGCGTGGACCTGGAGAGAAAGGTCGAATCACTTTTAGATGAGATAAATTTTCTGAGGAAGGTGCATGACGAAGAGGTGACCGAGTTGACTAATATGATCCAGGCGGCACAGATATCTGTGGAAGTGGAGTTGTCCAAGCCTGATCTGACCTCGGCCCTCAAGGACATTCGCGGCCAGTATGAGMutation Area - Mutant DNA Sequence
GATCTTGTTCTTAACAGTCACTACCAGAGGGAATATGACAGGCTGCTAAATGGGACAGTCCGCCGGGTATATAATGGCAAGCACACCAGTTTTAGACTCCCTTAGTTGTGACGAAGATGAGTTACGGATCAGACCACTAAGTCATCGAGGAAGACTTGCAGAAACTTCGAGGCAAGTTTGAGGAGGAGATCCGAGCGCGCGAGGAGGCAGAGCAGACGCTCAGATCCTACAAGAAAGATGTGGATGATGCCACCATGGTCCGCGTAGACCTGGAGAGAAAGGTCGAATCACTTTTAGATGAGATCAATTTTCTGAGGAAGGTGCATGACGAAGAGGTGACCGAGTTGACTAATATGATCCAGGCGGCACAGATATCTGTGGAAGTGGAGTTGTCCAAGCCTGATCTGACCTCGGCCCTCAAGGACATTCGCGGCCAGTATGAGGuide RNA Target Sites
TGTAGGATGAAGACATGTAGTGG
GCCGGTCCATTTCGACAGCGTGG
CATTGAACGCGACAGCATCGAGG
TGATGCCACCATGGTCCGCGTGG
Genotyping Primers
f, gatcttgtcttaacagtcactaccagaggg
r, CTCATACTGGCCGCGAATGTCC
wt_f, CTCTATCAGCAGGAGATCCGGGA
All three primers can be mixed together.
Alignment of WT and Mutant Sequences w/ gRNA Targets (cyan) and Genotyping Primers (pink) Shown on WT Sequence
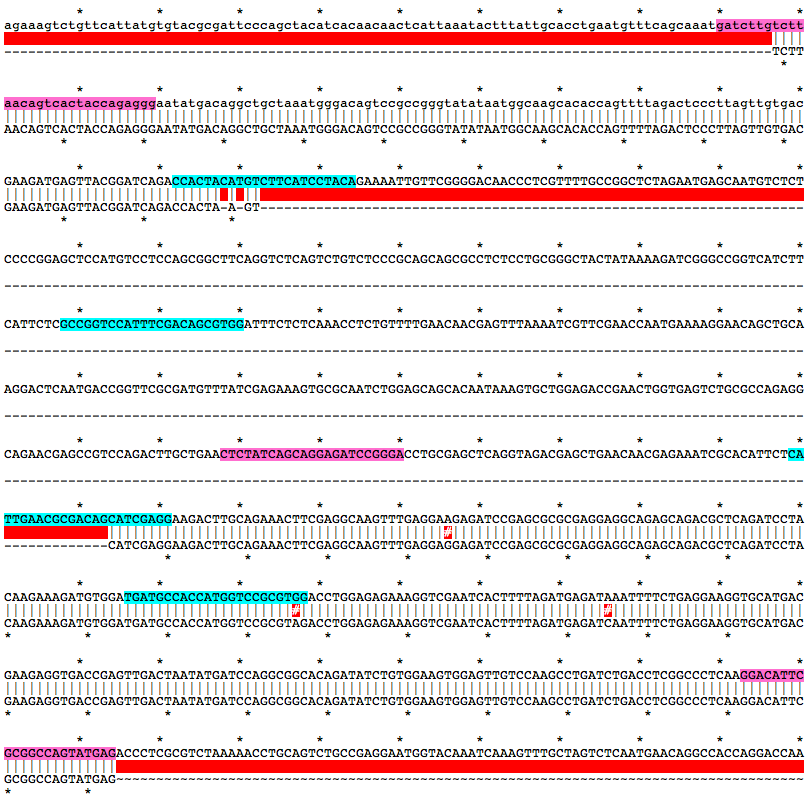

Allele
482bpDWT Genotyping Size
387WT Protein Sequence
MSYGSDHYMSSSYRKLFGDNPRFAGSRMSNVSPRSSMSSSGFRSQSVSRSSASPAGYYKR
SGRSSSFSPVHFDSVDFSQTSVLNNEFKIVRTNEKEQLQGLNDRFAMFIEKVRNLEQHNK
VLETELVSLRQRQNEPSRLAELYQQEIRDLRAQVDELNNEKSHILIERDSIEEDLQKLRG
KFEEEIRAREEAEQTLRSYKKDVDDATMVRVDLERKVESLLDEINFLRKVHDEEVTELTN
MIQAAQISVEVELSKPDLTSALKDIRGQYETLASKNLQSAEEWYKSKFASLNEQATRTNE
AMRATREEVNDYRRQLQSKTIEIETLRGTNESLERQIREMEEAHNAEVAGYQETIGQLDL
ELRNTKSEMARHLREYQDLLNVKMALDIEIAAYRKLLEGEETHFSSGVTFSSTPSITYGY
QSRSAFTSTRNPKKEREEEGHPKSKTAAKREENTDENVGIKKTEKNDAVDVNSN-
Mutant Protein Sequence
MSYGSDH-Mutant Genotyping Size
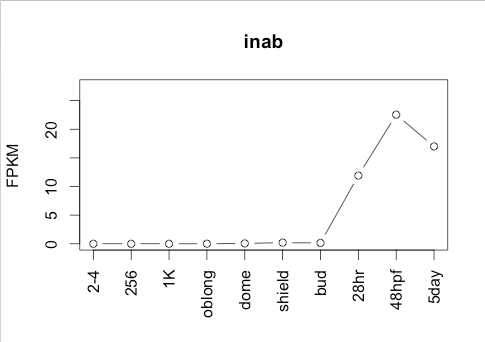
443Ribosome Profiling Development
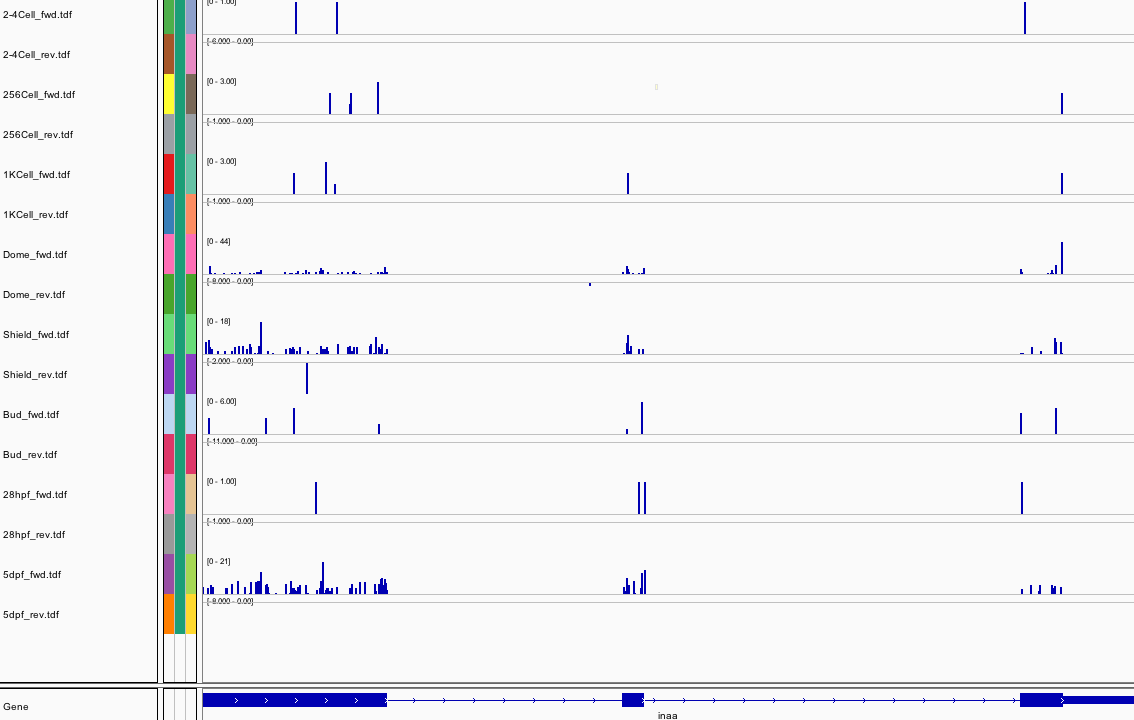
Transcript Plot
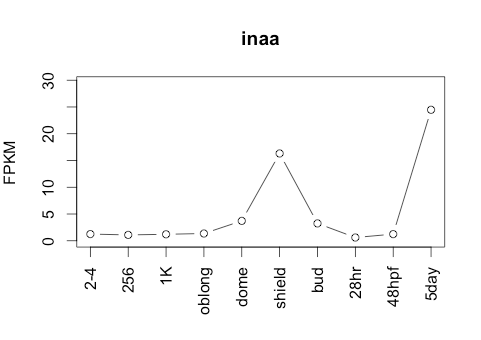
Sequence 2
Ensembl Gene Name
inabHarvard Allele
a286Mutation Area - WT DNA Sequence
tgcagtgaggaaggttgatgcgaggagaagcgtggctaaattattgattgggacggtcctgctataaaaggagacgccgccggagttctgctccttatcgtcaaacgccaagATGAGCTACGGATCCGACATCTACTCTGCCTCTTCCTACCGGAAGATCTTCGGAGATTCCACCCGTTTCTCAGCCTCTCCATCCCGGCTGAGCAGCTCCCGGAGCGGGTTTAAATCCCAGTCTGTGAACCGCCCCAACATCCCAAGCTCCTACAAGCGCTCCACCCGATCTGGCTTCCCGTCTTCTTCTCTGGACAGCTTCGATTTCACCCAGAGTACCGTGCTTAACAATGAGTTTAAAATCATCCGCACCAATGAGAAGGAGCAGATGCAAGGGCTCAACGACCGTTTCGCGATGTTCATTGAGAAAGTTCGCAATCTGGAGCAGCACAATAAAGTGCTGGAAACCGAACTCGTGACTCTGCGCCAGCGCCAGACAGAACCGTCCCGTCTGGCCGAACTTTACCAGCAGGAGATCCGCGAGCTGCGCTCCCAACTCGAGGAACTCAACGCCGAGAAGAACCAGATGATGTTCGAGCGCGACAGTATCGAGGAAGACCTACAGAAACTTCACGAGAAGTTTGAGGAAGAGATGAGAATGCGCGAGGAGGCCGAGCAAACCCTGAGAGCTTTCAAGAAGGACGTGGACAACGCCTCCATGGTGCGTCTAGACCTGGAGAAAAAGGTCGAATCCCTTCTGGATGAAATCAACTTTTTAAGGAAGGTGCACGAGGAGGAGGTGGCCGAGCTCATGAACATGATCCAGGCTAGCCAGGTGTCCGTGGAGATGGAAGTGGCCAAACCCGACCTCACCTCCGCCCTAAAGGATATTCGCGGCCAGTATGAAGCAATGGCGAGCAAGAACTTGCAGTCGGCTGAAGAGTGGTACAAGTCCAAGTTCGCCGACCTTAGCGAACAGGCCAACAAGAGCAACGAGGTCATTCGCGCTAGCAGGGAGGAGCTCAACGAGTTCAGGAGGCAGCTGCAGTCCAAGACCATCGAGATCGAGAGTCTGAGGGGTACAAATGMutation Area - Mutant DNA Sequence
tgcagtgaggaaggttgatgcgaggagaagcgtggctaaattattgattgggacggtcctgctataaaaggagacgccgccggagttctgctccttatcgtcaaacgccaagATGAGCTACGGATCCGACATCTACTCTGCCTCTTCCTACCGGAAGATCTTCGGAGATTCCACCCGTTTCTCAGCCTCTCCATCCCGGCTGAGCAGCTCCCGGAGCGGGTTTAAATCCCAGTCTGTGAACCGCCCAAGCTCCTACAAGCGCTCCACCCGATCTGGCTTCCCGTCTTCTTCTCTGGACAGCTTCGATTTCACCCAGAGTACCGTGCTTAACAATGAGTTTAAAATCATCCGCACCAATGAGAAGGAGCAGATGCAAGGGCTCAACGACCGTTTCGCGATGTTCATTGAGAAAGTTCGCAATCTGGAGCAGCACAATAAAGTGCTGGAAACCGAACTCGTGACTCTGCGCCAGCGCCAGACAGAACCGTCCCGTCTGGCCGAACTTTACCAGCAGGAGATCCGCGAGCTGCGCTCCCAACTCGAGGAACTCAACGCCGAGAAGAACCAGATGATGTTCGAGCGCGACAGTATCGAGGAAGACCTACAGAAACTTCACGAGAAGTTTGAGGAAGAGATGAGAATAAGACGGTAAACAAAGAGAAGGCAGTGAGGGAGAAGGTGTTAAAGTGAACGGTGCAAGGAGGAGGCCGAGCAAACCCTGAGAGCTTTCAAGAAGGACGTGGACAACGCCTCCATGGTGCGTCTAGACCTGGAGAAAAAGGTCGAATCCCTTCTGGATGAAATCAACTTTTTAAGGAAGGTGCACGAGGAGGAGGTGGCCGAGCTCATGAACATGATCCAGGCTAGCCAGGTGTCCGTGGAGATGGAAGTGGCCAAACCCGACCTCACCTCCGCCCTAAAGGATATTCGCGGCCAGTATGAAGCAATGGCGAGCAAGAACTTGCAGTCGGCTGAAGAGTGGTACAAGTCCAAGTTCGCCGACCTTAGCGAACAGGCCAACAAGAGCAACGAGGTCATTCGCGCTAGCAGGGAGGAGCTCAACGAGTTCAGGAGGCAGCTGCAGTCCAAGACCATCGAGATCGAGAGTCTGAGGGGTACAAATGGuide RNA Target Sites
GAAGATCTTCCGGTAGGAAGAGG
AGGAGCTTGGGATGTTGGGGCGG
GGAAGAGATGAGAATGCGCGAGG
GCGAATATCCTTTAGGGCGGAGG
Genotyping Primers
f, GCGCTCCCAACTCGAGGAACTCA
r, ACCTCCTCCTCGTGCACCTTCCT
Alignment of WT and Mutant Sequences w/ gRNA Targets (cyan) and Genotyping Primers (pink) Shown on WT Sequence

Allele
9bpD,54bpIWT Genotyping Size
255WT Protein Sequence
MSYGSDIYSASSYRKIFGDSTRFSASPSRLSSSRSGFKSQSVNRPNIPSSYKRSTRSGFP
SSSLDSFDFTQSTVLNNEFKIIRTNEKEQMQGLNDRFAMFIEKVRNLEQHNKVLETELVT
LRQRQTEPSRLAELYQQEIRELRSQLEELNAEKNQMMFERDSIEEDLQKLHEKFEEEMRM
REEAEQTLRAFKKDVDNASMVRLDLEKKVESLLDEINFLRKVHEEEVAELMNMIQASQVS
VEMEVAKPDLTSALKDIRGQYEAMASKNLQSAEEWYKSKFADLSEQANKSNEVIRASREE
LNEFRRQLQSKTIEIESLRGTNESLERQIHEMEDRHNAEIMGYQDSIGELENDLRTTKSE
MARHLREYQDLLNVKMALDIEIAAYRKLLEGEETRISTGITYPTPASVSSYSYQSRLYSS
TSMSSKKEVKDDDDKQQSGKPGKGSSQSDDSKKNDKIDSGDLNSTNQKN-
Mutant Protein Sequence
MSYGSDIYSASSYRKIFGDSTRFSASPSRLSSSRSGFKSQSVNRPSSYKRSTRSGFPSSS
LDSFDFTQSTVLNNEFKIIRTNEKEQMQGLNDRFAMFIEKVRNLEQHNKVLETELVTLRQ
RQTEPSRLAELYQQEIRELRSQLEELNAEKNQMMFERDSIEEDLQKLHEKFEEEMRIRR-
Mutant Genotyping Size
309Ribosome Profiling Development
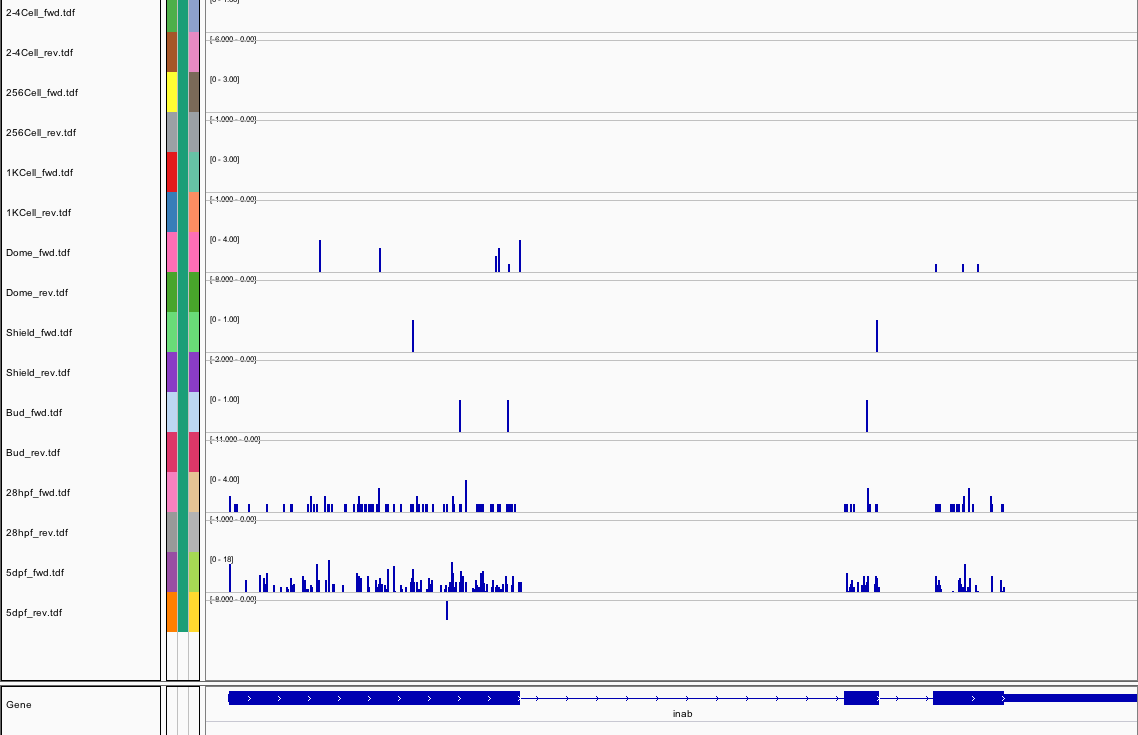
Transcript Plot
Behavior Data
Behavior Data Summary
NoneBehavior Data Description 1
Dark flash block 1 start Merged section Pvalue = not significant 26 hethet vs 18 homhom ribgraph_mean_ribbon_fullboutdata_dpix_a0darkflash103.pngBehavior Data Graph 1
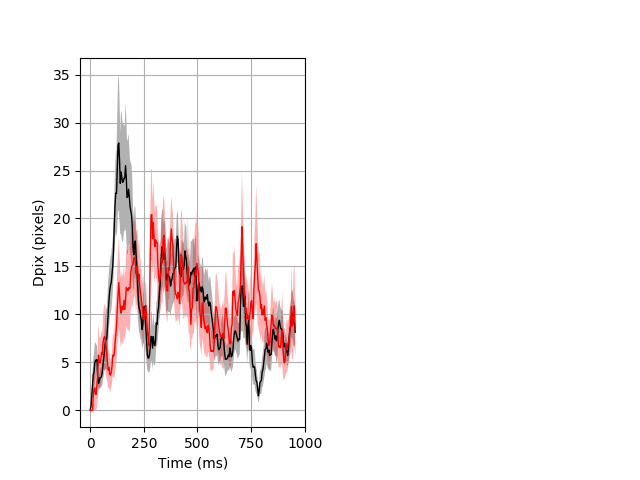
Behavior Data Description 2
Day all prepulse tap Merged section Pvalue = not significant 26 hethet vs 18 homhom ribgraph_mean_ribbon_fullboutdata_dpix_dayprepulseinhibition100d.pngBehavior Data Graph 2
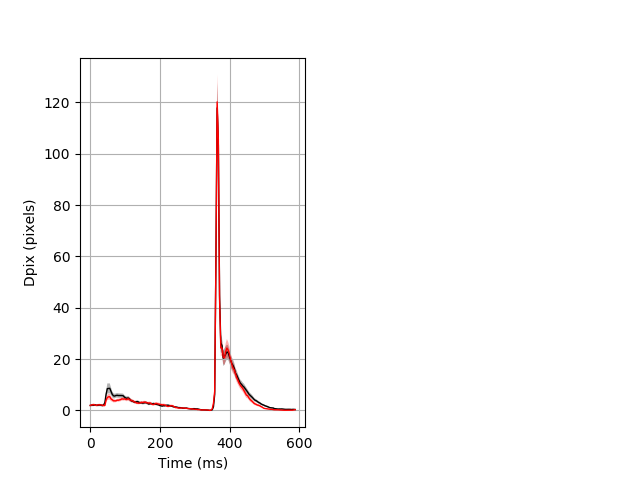
Behavior Data Description 3
Features of movement entire protocol Graph Pvalue = not significant Merged section Pvalue = not significant 26 hethet vs 18 homhom ribgraph_mean_ribbonbout_aveboutdisp_10min_combo.pngBehavior Data Graph 3
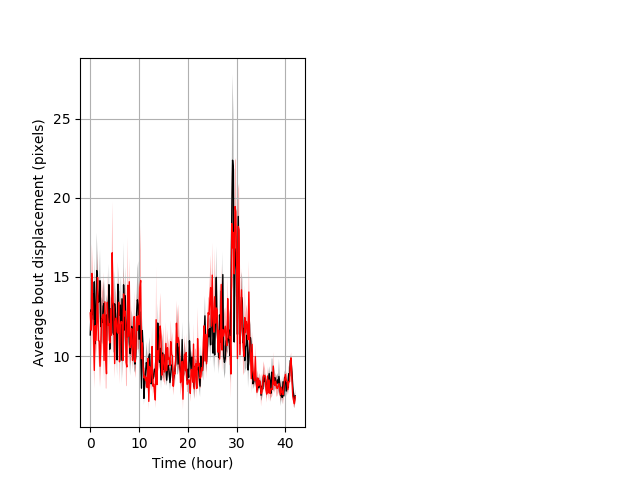
Behavior Data Description 4
Frequency of movement entire protocol Graph Pvalue = not significant Merged section Pvalue = not significant 26 hethet vs 18 homhom ribgraph_mean_ribbonbout_dpixnumberofbouts_10min_combo.pngBehavior Data Graph 4
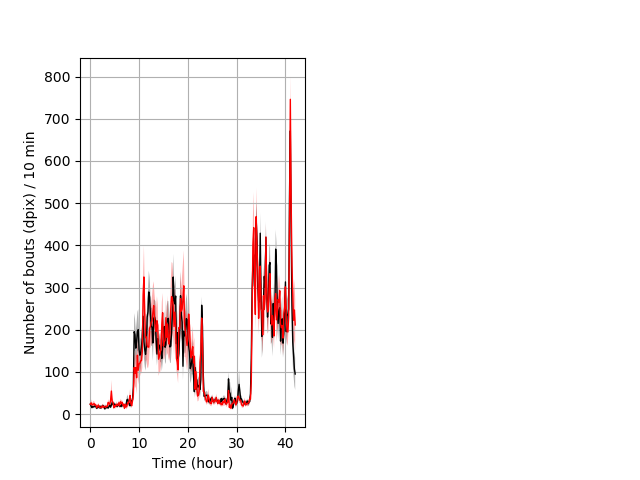
Behavior Data Description 5
Location in well entire protocol Graph Pvalue = not significant Merged section Pvalue = not significant 26 hethet vs 18 homhom ribgraph_mean_ribbonbout_averhofrac_10min_combo.pngBehavior Data Graph 5
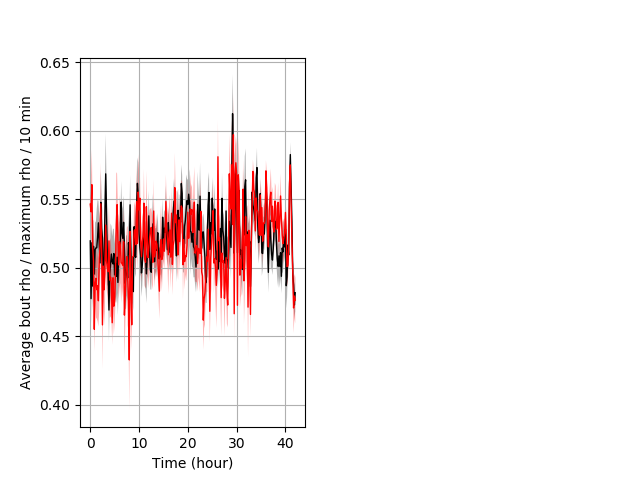
Behavior Data Description 6
NoneBehavior Data Graph 6
NoneBehavior Data Description 7
NoneBehavior Data Graph 7
NoneBehavior Data Description 8
NoneBehavior Data Graph 8
NoneBehavior Data Description 9
NoneBehavior Data Graph 9
NoneBehavior Data Description 10
NoneBehavior Data Graph 10
NoneBehavior Data Description 11
NoneBehavior Data Graph 11
NoneBehavior Data Description 12
NoneBehavior Data Graph 12
NoneBehavior Data Description 13
NoneBehavior Data Graph 13
NoneBehavior Data Description 14
NoneBehavior Data Graph 14
None