Locus Rank
3Gene
Id
9Gene Name
CYP17A1Duplicated
FalseMaternal
TrueGene Not Within Locus (Nearby
FalseGene Description
This gene encodes a member of the cytochrome P450 superfamily of enzymes. The cytochrome P450 proteins are monooxygenases which catalyze many reactions involved in drug metabolism and synthesis of cholesterol, steroids and other lipids. This protein localizes to the endoplasmic reticulum. It has both 17alpha-hydroxylase and 17,20-lyase activities and is a key enzyme in the steroidogenic pathway that produces progestins, mineralocorticoids, glucocorticoids, androgens, and estrogens. Mutations in this gene are associated with isolated steroid-17 alpha-hydroxylase deficiency, 17-alpMouse Phenotype
Homozygous null embryos display early embryonic lethality.Additional Image
Allen Brain
Allen Regions
Zfin In Situ
Zfin Brain Areas
Published Zebrafish Pehnotype
Not In Allen Brain
TrueZFIN Link
http://zfin.org/ZDB-GENE-040213-2Allen Link
http://mouse.brain-map.org/gene/show/12855More Additional Images
Papers
http://www.ncbi.nlm.nih.gov/pubmed/22800812
http://www.ncbi.nlm.nih.gov/pubmed/21692878
http://www.sciencedirect.com/science/article/pii/S0016648011003649
Locus Rank
Locus Rank
3Genes in Region
ARL3, AS3MT, C10orf32, CNNM2, CYP17A1, INA, NT5C2, PCGF6, PDCD11, SFXN2, TAF5, TRIM8, USMG5, WBP1LAssociated Snps
rs11191419 (shown), chr10_104957618_I, rs7907645, rs55833108GWAS Region (hg19)
chr10:104423800-105165583Genes Skipping
AS3MT, NT5C2, PCGF6, PDCD11, SFXN2, TAF5, TRIM8, USMG5, WBP1LRicopili Plot Surrounding Area
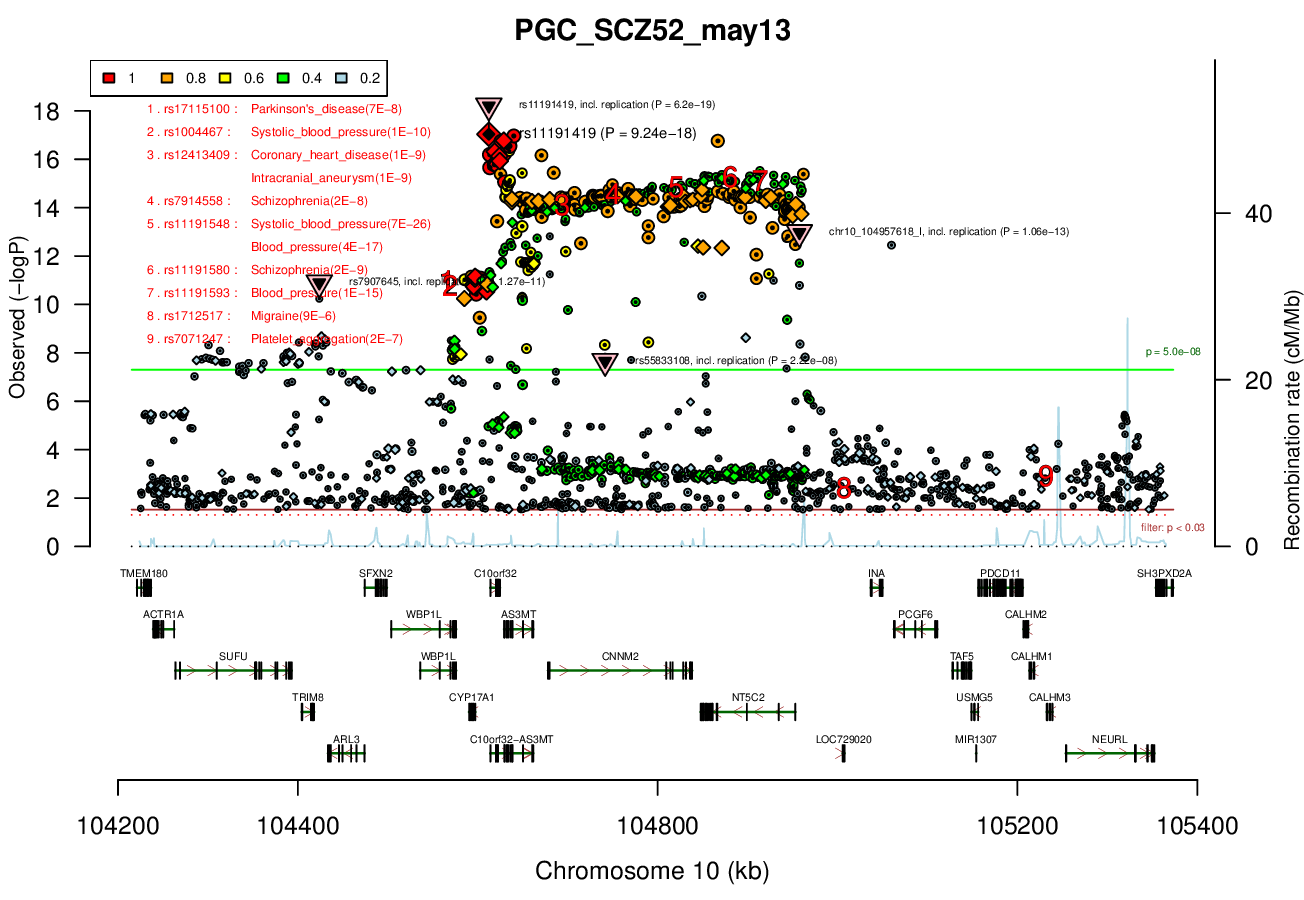
Distance On Each Side Of Locus In Above Plot
0.2 MBRicopili Plot
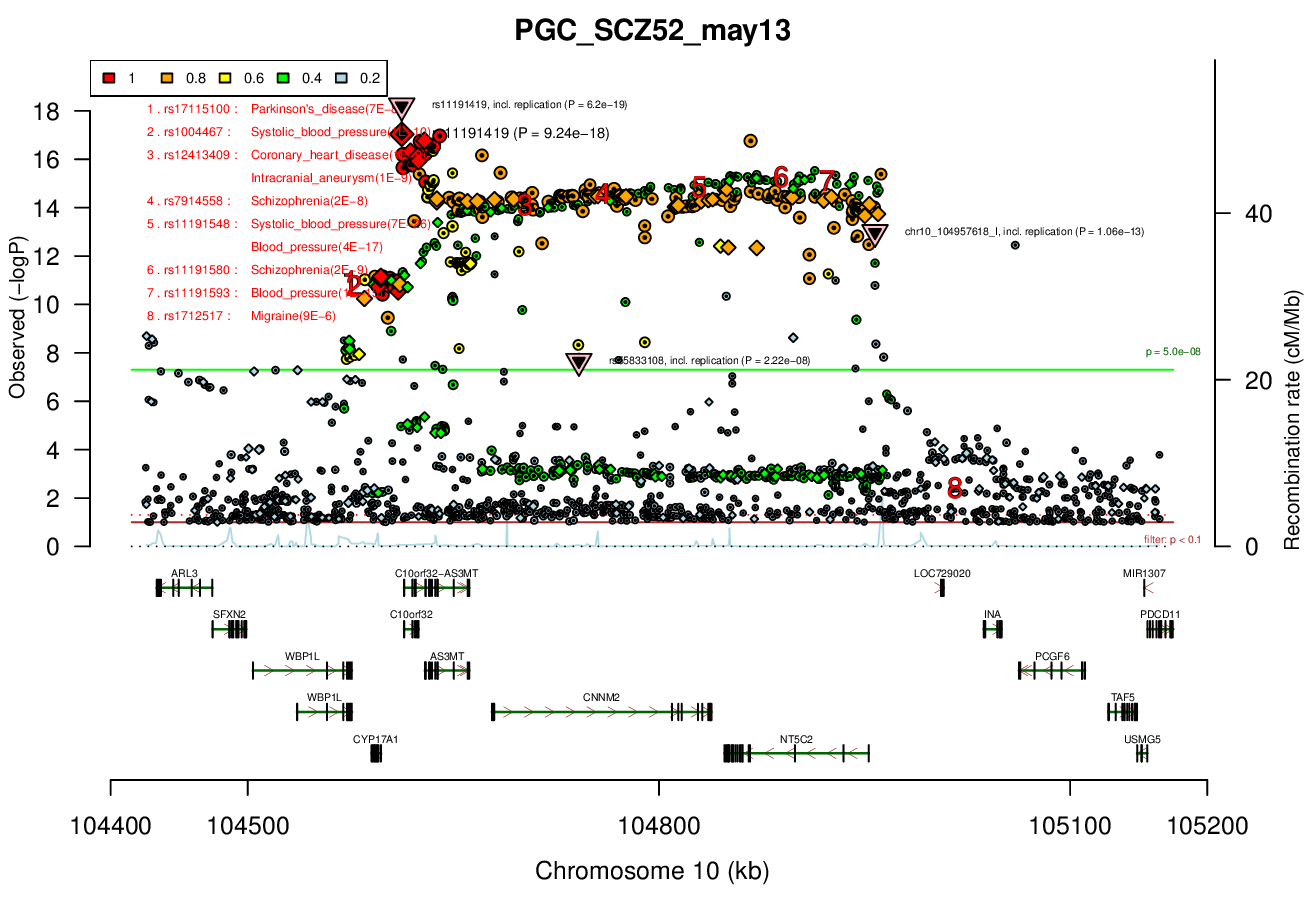
Gwas And Surrounding Region Pic
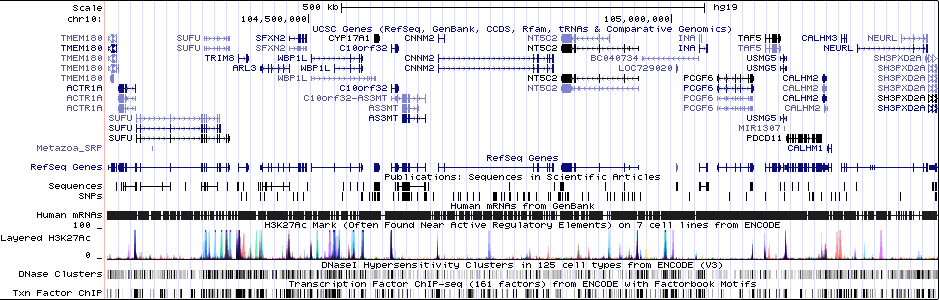
Gwas Region Pic
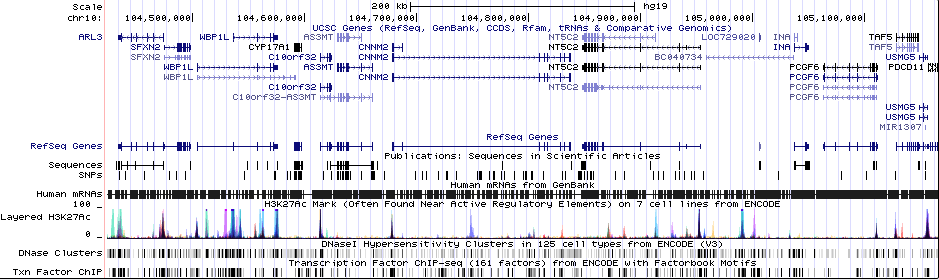
Sequence 1
Ensembl Gene Name
cyp17a1Harvard Allele
a259Mutation Area - WT DNA Sequence
CTGGAGGACACTCAGTTGAATATAGCTGACAATGGCTGAAGCACTCATCCTGCCCTGGCTGCTCTGTTTAAGCCTGTTCTCCGCAGTGACTCTGGCAGCTCTGTATCTCAAACAGAAGATGAATGGATTTGTGCCAGCAGGAAATAGATCTCCTCCAAGTCTCCCTTCACTGCCCATCATCGGGAGTCTGATGAGCCTGGTGAGCGACAGTCCTCCGCACATCTTCTTTCAGGACCTGCAGAAGAAATACGGAGATCTGTATTCCCTCATGATGGGCTCCCACAAACTGCTCATTGTGAACAACCACCATCATGCGAAGGAGATCCTGATCAAAAAAGGAAAAATATTTGCAGGGAGGCCACGGACTgtaagtacatacaatcacattcagtttaattagccatttttagctgtttaaattttgttaatttaaatatggcccatctgcatatggttaaactaaatttacMutation Area - Mutant DNA Sequence
CTGGAGGACACTCAGTTGAATATAGCTGACAATGGCTGAAGCACTCATCCTGCCCTGGCTGCTCTGTTTAAGCCTGTTCTCAGCAGTGACTCTGGCAGCTCTGTATCTCAAACAGAAGATGAATGGATTTGTGCCAGCAGGAAACAGATCTCCTCGAAGGAGATCCTGATCAAAAAAGGAAAAATATTTGCAGGGAGGCCACGGACTGTAAGTACATACAATCACATTCAGTTTAATTAGCCATTTTTAGCTGTTTAAATTTTGTTAATTTAAATATGGCCCATCTGCATATGGTTAAACTAAATTTACGuide RNA Target Sites
GGCAGTGAAGGGAGACTTGGAGG
GCGGAGGACTGTCGCTCACCAGG
GTTTGTGGGAGCCCATCATGAGG
GGATCTCCTTCGCATGATGGTGG
Genotyping Primers
f, CTGGAGGACACTCAGTTGAATATAGC
r,gtaaatttagtttaaccatatgcagatgggcc
Alignment of WT and Mutant Sequences w/ gRNA Targets (cyan) and Genotyping Primers (pink) Shown on WT Sequence
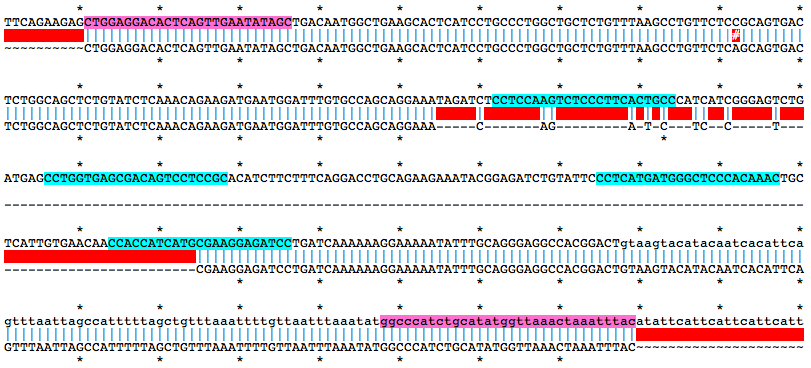
Allele
160bpDWT Genotyping Size
469WT Protein Sequence
MAEALILPWLLCLSLFSAVTLAALYLKQKMNGFVPAGNRSPPSLPSLPIIGSLMSLVSDS
PPHIFFQDLQKKYGDLYSLMMGSHKLLIVNNHHHAKEILIKKGKIFAGRPRTVTTDLLTR
DGKDIAFADYSSTWKFHRKMVHGALCMFGEGSVSIEKIICREASSMCEVLTESQNSAVDL
GPELTRAVTNVVCALCFNSSYKRGDAEFESMLQYSQGIVDTVAKDSLVDIFPWLQIFPNK
DLRILRQCISIRDKLLQKKYEEHKVTYSDNVQRDLLDALLRAKRSSENNNSSTRDVGLTE
DHVLMTVGDIFGAGVETTTTVLKWSIAYLVHNPQVQRKIQEELDSKIGKERHPQLSDRGN
LPYLEATIREVLRIRPVSPLLIPHVALQDSSVGEYTVQKGTRVVINLWSLHHDEKEWKNP
ELFDPGRFLNEEGDGLCCPSGSYLPFGAGVRVCLGEALAKMELFLFLAWILQRFTLEMPT
GQPLPDLQGKFGVVLQPKKFKVVAKVRADWEKSPLMQHC-
Mutant Protein Sequence
MAEALILPWLLCLSLFSAVTLAALYLKQKMNGFVPAGNRSPRRRS-Mutant Genotyping Size
309Ribosome Profiling Development
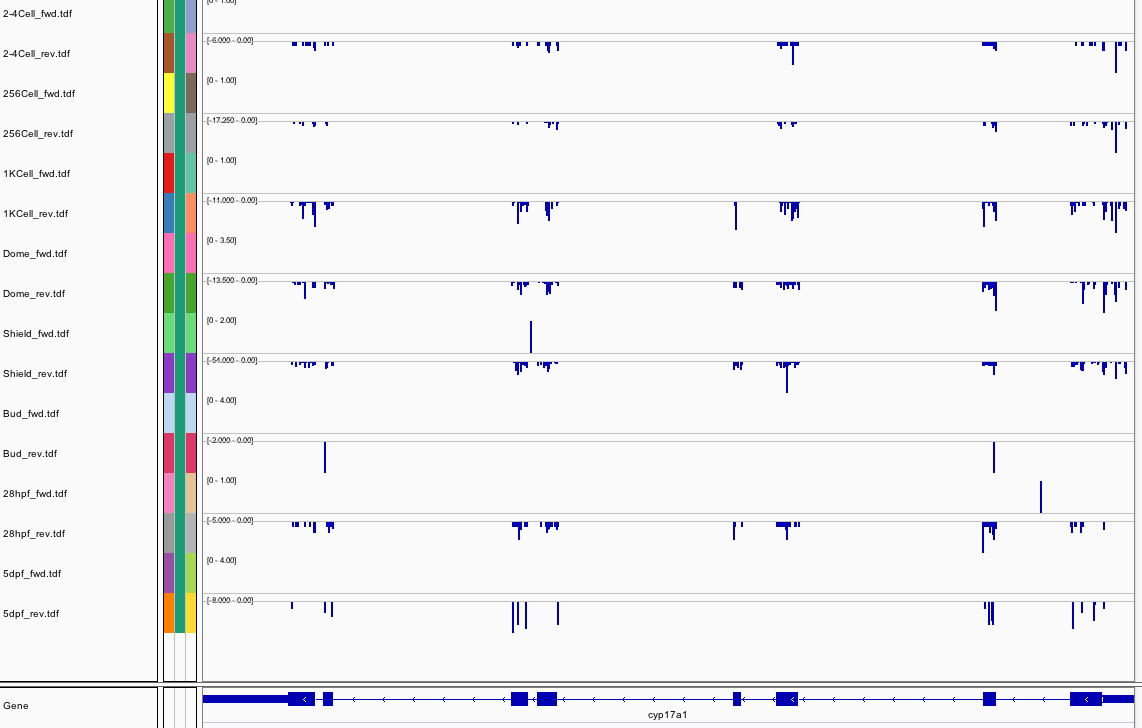
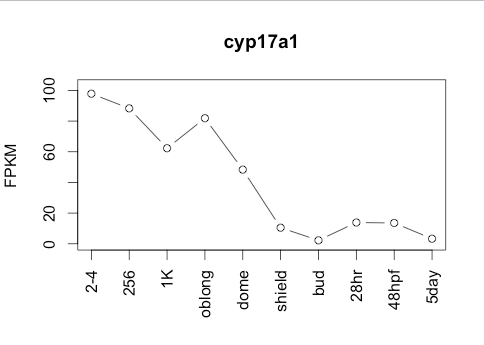
Transcript Plot
Behavior Data
Behavior Data Summary
NoneBehavior Data Description 1
Dark flash block 1 start Merged section Pvalue = 0.0231 9 wt vs 9 hom ribgraph_mean_ribbon_fullboutdata_dpix_a0darkflash103.pngBehavior Data Graph 1
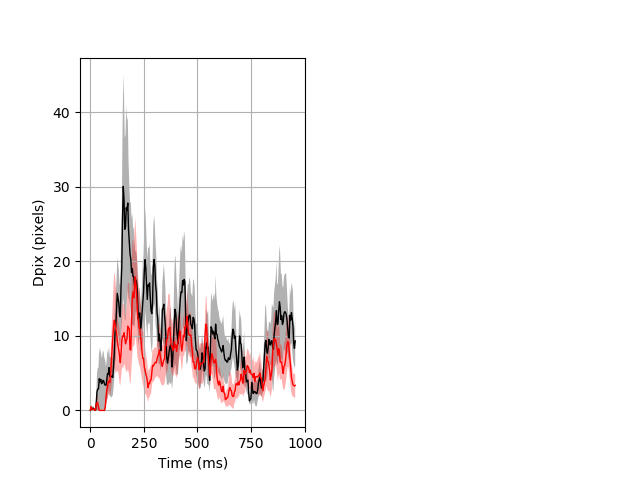
Behavior Data Description 2
Day all prepulse tap Merged section Pvalue = not significant 9 wt vs 9 hom ribgraph_mean_ribbon_fullboutdata_dpix_dayprepulseinhibition100d.pngBehavior Data Graph 2
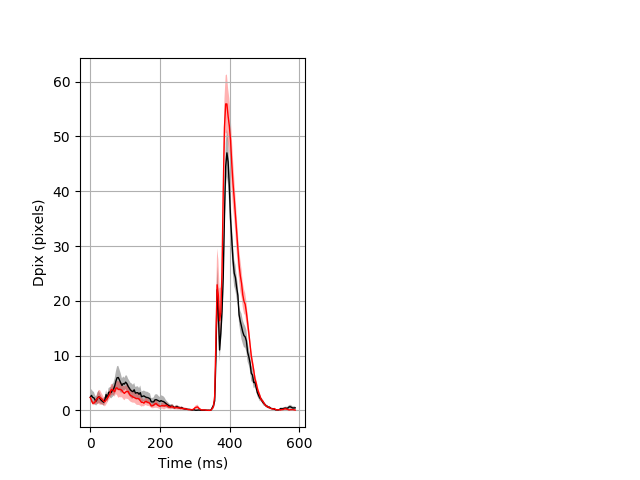
Behavior Data Description 3
Features of movement entire protocol Graph Pvalue = not significant Merged section Pvalue = not significant 9 wt vs 9 hom ribgraph_mean_ribbonbout_aveboutdisp_10min_combo.pngBehavior Data Graph 3
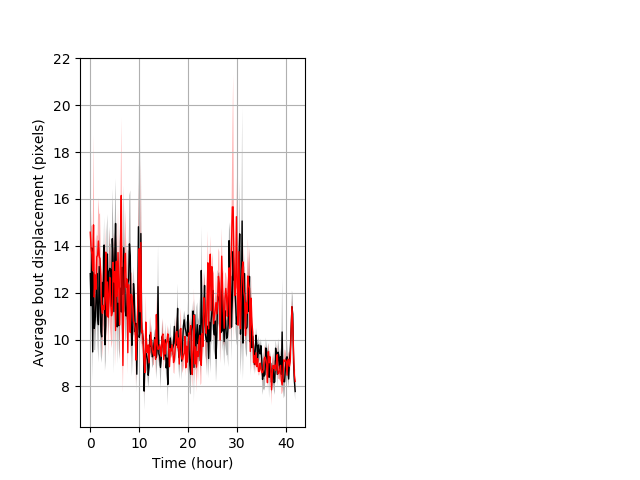
Behavior Data Description 4
Frequency of movement entire protocol Graph Pvalue = not significant Merged section Pvalue = not significant 9 wt vs 9 hom ribgraph_mean_ribbonbout_dpixnumberofbouts_10min_combo.pngBehavior Data Graph 4
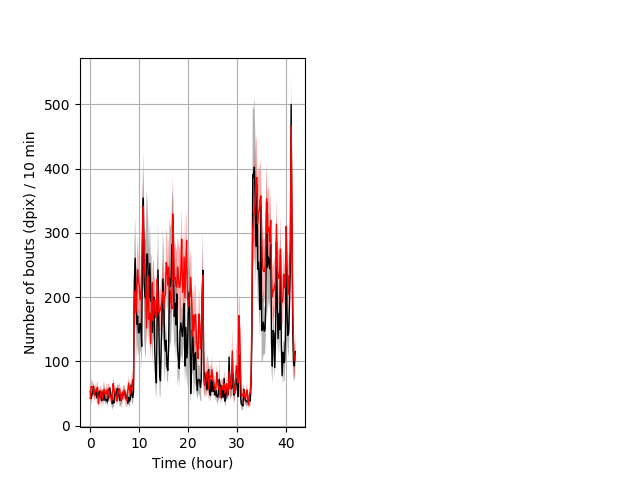
Behavior Data Description 5
Location in well entire protocol Graph Pvalue = not significant Merged section Pvalue = not significant 9 wt vs 9 hom ribgraph_mean_ribbonbout_averhofrac_10min_combo.pngBehavior Data Graph 5
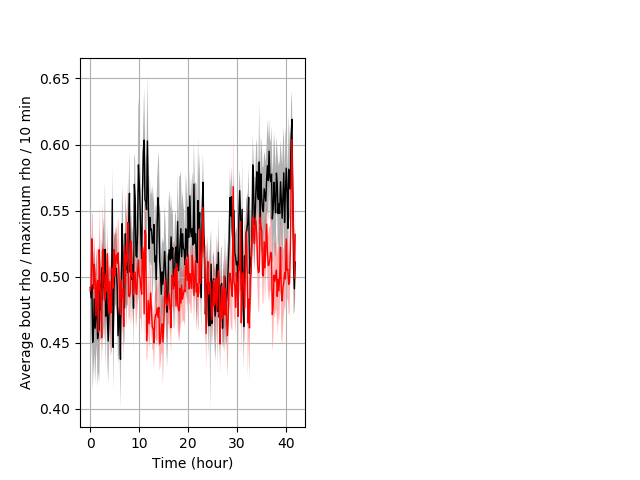
Behavior Data Description 6
NoneBehavior Data Graph 6
NoneBehavior Data Description 7
NoneBehavior Data Graph 7
NoneBehavior Data Description 8
NoneBehavior Data Graph 8
NoneBehavior Data Description 9
NoneBehavior Data Graph 9
NoneBehavior Data Description 10
NoneBehavior Data Graph 10
NoneBehavior Data Description 11
NoneBehavior Data Graph 11
NoneBehavior Data Description 12
NoneBehavior Data Graph 12
NoneBehavior Data Description 13
NoneBehavior Data Graph 13
NoneBehavior Data Description 14
NoneBehavior Data Graph 14
None